

Community Blog
Keep up-to-date on postgraduate related issues with our quick reads written by students, postdocs, professors and industry leaders.
Types of Research – Explained with Examples
- By DiscoverPhDs
- October 2, 2020

Types of Research
Research is about using established methods to investigate a problem or question in detail with the aim of generating new knowledge about it.
It is a vital tool for scientific advancement because it allows researchers to prove or refute hypotheses based on clearly defined parameters, environments and assumptions. Due to this, it enables us to confidently contribute to knowledge as it allows research to be verified and replicated.
Knowing the types of research and what each of them focuses on will allow you to better plan your project, utilises the most appropriate methodologies and techniques and better communicate your findings to other researchers and supervisors.
Classification of Types of Research
There are various types of research that are classified according to their objective, depth of study, analysed data, time required to study the phenomenon and other factors. It’s important to note that a research project will not be limited to one type of research, but will likely use several.
According to its Purpose
Theoretical research.
Theoretical research, also referred to as pure or basic research, focuses on generating knowledge , regardless of its practical application. Here, data collection is used to generate new general concepts for a better understanding of a particular field or to answer a theoretical research question.
Results of this kind are usually oriented towards the formulation of theories and are usually based on documentary analysis, the development of mathematical formulas and the reflection of high-level researchers.
Applied Research
Here, the goal is to find strategies that can be used to address a specific research problem. Applied research draws on theory to generate practical scientific knowledge, and its use is very common in STEM fields such as engineering, computer science and medicine.
This type of research is subdivided into two types:
- Technological applied research : looks towards improving efficiency in a particular productive sector through the improvement of processes or machinery related to said productive processes.
- Scientific applied research : has predictive purposes. Through this type of research design, we can measure certain variables to predict behaviours useful to the goods and services sector, such as consumption patterns and viability of commercial projects.

According to your Depth of Scope
Exploratory research.
Exploratory research is used for the preliminary investigation of a subject that is not yet well understood or sufficiently researched. It serves to establish a frame of reference and a hypothesis from which an in-depth study can be developed that will enable conclusive results to be generated.
Because exploratory research is based on the study of little-studied phenomena, it relies less on theory and more on the collection of data to identify patterns that explain these phenomena.
Descriptive Research
The primary objective of descriptive research is to define the characteristics of a particular phenomenon without necessarily investigating the causes that produce it.
In this type of research, the researcher must take particular care not to intervene in the observed object or phenomenon, as its behaviour may change if an external factor is involved.
Explanatory Research
Explanatory research is the most common type of research method and is responsible for establishing cause-and-effect relationships that allow generalisations to be extended to similar realities. It is closely related to descriptive research, although it provides additional information about the observed object and its interactions with the environment.
Correlational Research
The purpose of this type of scientific research is to identify the relationship between two or more variables. A correlational study aims to determine whether a variable changes, how much the other elements of the observed system change.
According to the Type of Data Used
Qualitative research.
Qualitative methods are often used in the social sciences to collect, compare and interpret information, has a linguistic-semiotic basis and is used in techniques such as discourse analysis, interviews, surveys, records and participant observations.
In order to use statistical methods to validate their results, the observations collected must be evaluated numerically. Qualitative research, however, tends to be subjective, since not all data can be fully controlled. Therefore, this type of research design is better suited to extracting meaning from an event or phenomenon (the ‘why’) than its cause (the ‘how’).
Quantitative Research
Quantitative research study delves into a phenomena through quantitative data collection and using mathematical, statistical and computer-aided tools to measure them . This allows generalised conclusions to be projected over time.

According to the Degree of Manipulation of Variables
Experimental research.
It is about designing or replicating a phenomenon whose variables are manipulated under strictly controlled conditions in order to identify or discover its effect on another independent variable or object. The phenomenon to be studied is measured through study and control groups, and according to the guidelines of the scientific method.
Non-Experimental Research
Also known as an observational study, it focuses on the analysis of a phenomenon in its natural context. As such, the researcher does not intervene directly, but limits their involvement to measuring the variables required for the study. Due to its observational nature, it is often used in descriptive research.
Quasi-Experimental Research
It controls only some variables of the phenomenon under investigation and is therefore not entirely experimental. In this case, the study and the focus group cannot be randomly selected, but are chosen from existing groups or populations . This is to ensure the collected data is relevant and that the knowledge, perspectives and opinions of the population can be incorporated into the study.
According to the Type of Inference
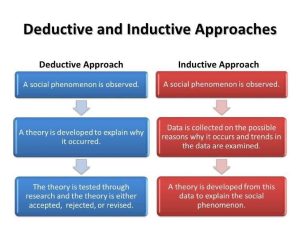
Deductive investigation.
In this type of research, reality is explained by general laws that point to certain conclusions; conclusions are expected to be part of the premise of the research problem and considered correct if the premise is valid and the inductive method is applied correctly.
Inductive Research
In this type of research, knowledge is generated from an observation to achieve a generalisation. It is based on the collection of specific data to develop new theories.
Hypothetical-Deductive Investigation
It is based on observing reality to make a hypothesis, then use deduction to obtain a conclusion and finally verify or reject it through experience.

According to the Time in Which it is Carried Out
Longitudinal study (also referred to as diachronic research).
It is the monitoring of the same event, individual or group over a defined period of time. It aims to track changes in a number of variables and see how they evolve over time. It is often used in medical, psychological and social areas .
Cross-Sectional Study (also referred to as Synchronous Research)
Cross-sectional research design is used to observe phenomena, an individual or a group of research subjects at a given time.
According to The Sources of Information
Primary research.
This fundamental research type is defined by the fact that the data is collected directly from the source, that is, it consists of primary, first-hand information.
Secondary research
Unlike primary research, secondary research is developed with information from secondary sources, which are generally based on scientific literature and other documents compiled by another researcher.

According to How the Data is Obtained
Documentary (cabinet).
Documentary research, or secondary sources, is based on a systematic review of existing sources of information on a particular subject. This type of scientific research is commonly used when undertaking literature reviews or producing a case study.
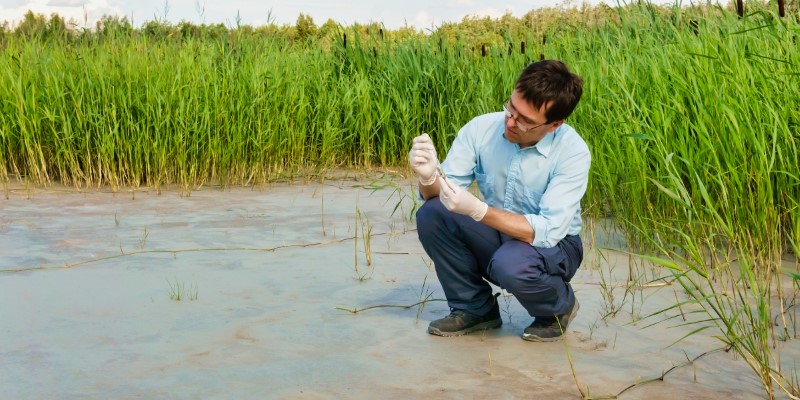
Field research study involves the direct collection of information at the location where the observed phenomenon occurs.
From Laboratory
Laboratory research is carried out in a controlled environment in order to isolate a dependent variable and establish its relationship with other variables through scientific methods.
Mixed-Method: Documentary, Field and/or Laboratory
Mixed research methodologies combine results from both secondary (documentary) sources and primary sources through field or laboratory research.

Reference management software solutions offer a powerful way for you to track and manage your academic references. Read our blog post to learn more about what they are and how to use them.

Starting your PhD can feel like a daunting, exciting and special time. They’ll be so much to think about – here are a few tips to help you get started.

The scope of the study is defined at the start of the study. It is used by researchers to set the boundaries and limitations within which the research study will be performed.
Join thousands of other students and stay up to date with the latest PhD programmes, funding opportunities and advice.

Browse PhDs Now

The unit of analysis refers to the main parameter that you’re investigating in your research project or study.

The term monotonic relationship is a statistical definition that is used to describe the link between two variables.

Dr Joseph gained her PhD in Chemistry from the University of Cambridge in 2018 and is now a Postdoctoral Research Associate in Physics at the University. Her research is on better understanding how cells organises its contents via the process of liquid-liquid phase separation.

Maria is a 1st year PhD student at the University of Birmingham, researching how to employ and exploit the biology of human gamma delta T cells for development of cancer immunotherapy.
Join Thousands of Students
- Find Studies
For Researchers
- Learn about Research
What are some different types of research studies?

There are many different types of research studies. Generally, there are two major types of studies available on Research for Me @UNC: research studies and clinical trials . When a research study is about disease or human health, it is called a clinical research study. When a research study involves drugs or other therapies that aim to slow or stop a disease, then it is called a clinical trial. Volunteers are an important part of all of these research studies! Explore other types of research studies below.
Survey - Survey studies ask people questions about their knowledge, attitudes, and feelings about a wide range of topics. You can complete these surveys online, over the phone, or by mail. Sometimes, these studies might also be in-person interviews or group discussions.
Lifestyle - Lifestyle studies look at what happens when people participate in different types of activities over a set period of time. You may attend activity sessions in a center or clinic or be asked to change the way that you do something in your daily activities. Often, these studies are interested in how changes in behavior can affect our health or other parts of our lives.
Drug - Drug studies are heavily regulated by the US Government. Studies often involve medications that are not currently available to the general public. They are called “investigational” drugs and have not yet been approved by the FDA (US Food and Drug Administration) for your normal health care provider to prescribe. Other drug studies may involve comparisons between two or more FDA-approved medications.
Device - Device studies are done to learn if a new medical device helps relieve a certain medical condition. Devices you may be familiar with are pacemakers, diabetes testing meters, and hearing aids. These studies usually involve devices that are not currently available to the general public and have not been approved for use by the FDA. Sometimes, they may be studying an FDA-approved device, but for use in treating a new condition.
Procedure - Procedure studies learn about the safety and effectiveness of certain medical procedures. Sometimes they compare a new medical procedure to one already in use. Procedures might include things like imaging (x-rays), stitches, blood tests, and surgeries.
Medical Outcomes - Outcomes research studies the end results (outcomes) of the structure and processes of the health care system on the health and well-being of patients and populations. These studies look at clinical practices to see if there are better ways for doctors to help patients manage their medical care. Outcomes research often considers patients’ experiences, preferences, and values – all of which may affect whether or not a medical treatment is best for them.
Community-based - Community-based research is done through a true partnership of community leaders and organizations with a UNC researcher or research team. The ideas are driven by community members and the research incorporates voices of all involved. These studies aim to understand problems impacting communities and contribute to solutions through policy or social change.
Copyright © 2013-2022 The NC TraCS Institute, the integrated home of the NIH Clinical and Translational Science Awards (CTSA) Program at UNC-CH. This website is made possible by CTSA Grant UL1TR002489 and the National Center for Advancing Translational Sciences.
Helpful Links
- My UNC Chart
- Find a Doctor
- Accessibility
- Privacy Policy

The Two Main Research Types – A Brief Overview
In this module, we will have a short overview of the two main types of research before we dive further into learning more about each in detail.
Content includes:
- Qualitative methods brief introductory overview
- Quantitative methods brief introductory overview
Objectives:
- Identify the three main types of qualitative research.
- Describe the processes of inductive versus deductive reasoning, and the types of research for both.
- Distinguish quantitative experimental and nonexperimental research.
Here we go! We are now going to start learning a bit about research. Remember, research is the underpinning for EBP. Research provides the evidence, and EBP takes that evidence and embeds it into practice to improve clinical outcomes.
The two major classes of research are:
- Qualitative Research – subjective, seeks a human’s experience as a narrative
- Quantitative Research – objective, seeks to statistically make inferences about a sample to generalize to the larger population
We need to have a solid understanding of the difference between the two main types of research before we study the nuances of each.
Three Main Types of Qualitative Research
Qualitative Research : Qualitative research is rooted in research that originated in anthropology, sociology, and psychology. Qualitative research is not experimental, it seeks to understand the lived experiences in humans and seeks to understanding meaning, and it is subjective in nature. The overarching goal of qualitative research is theory-generating. It is an inductive process (inductive reasoning). Most often, qualitative research features an interview style. This allows the researcher to ask open-ended questions and the participants share their experiences and/or explanation of particular meanings in life.
Qualitative research differs from quantitative research in that:
- It is completely subjective .
- It utilizes an inductive (versus deductive) approach.
- It does not utilize a hypothesis.
- It generates a theory from the data to explain the social phenomenon that the researchers were interested in.
- The researcher is involved with the participants for data collection.
- The data is analyzed with a thematic nature. That is, themes from the collected narratives are analyzed to see trends or themes in what the participants shared.
- The results are not generalizable to the population.
There are three types of qualitative research designs:
| Grounded Theory | This type of qualitative research seeks to understand and describe social psychological processes. | Example: |
| Phenomenology | This is concerned with the lived experiences of humans. | Example:
|
| Ethnography | This is concerned with learning about patterns and lifeways of cultural groups. Often these researchers go to the culture itself (fieldwork) to interview the participants in their natural settings. | Example:
|
Inductive versus Deductive Reasoning
The main difference between inductive and deductive reasoning is that inductive reasoning aims at developing a theory while deductive reasoning aims at testing an existing theory.
Think of inductive (theory producing) as to qualitative research and deductive (theory testing) as to quantitative research.
Inductive reasoning moves from specific observations to broad generalizations, and deductive reasoning the other way around.
Both approaches are used in various types of research, and it’s not uncommon to combine them in one large study.
Here is a qualitative study in which the researchers conducted interviews in order to obtain the subjective perspectives of the participants.

Quantitative Experimental and Nonexperimental Research.
Quantitative Research: In quantitative research, the goal is to utilize the statistical data to generalize results to the population studied. Some key features include utilizing the statistics to help answer the clinical question and determine whether the hypothesis is indeed statistically supported.
There are two main types of quantitative research:
- Experimental : In experimental research, the researcher introduces an intervention or treatment.
- Non-Experimental : In non-experimental research, the researcher does not introduce an intervention or treatment, but instead acts as a bystander. Meaning, they collect data without introducing a treatment.
We will explore those two types in much detail in the next module.
Quantitative research differs from qualitative research in that:
- It is completely objective .
- It utilizes a deductive (versus inductive) approach.
- It utilizes a hypothesis(es).
- It tests a theory.
- The researcher is usually not directly involved with the participants for data collection in order to minimize bias.
- The data is analyzed statistically in order to generalize results to the larger population.
Experimental Research : In the following article, the researchers introduced an intervention, which was a “Program for Enhancing the Positive Aspects of Caregiving” (a particular education program).

Non-experimental Research : In the following article, the researchers did not introduce an intervention or treatment. They handed out surveys for the participants to complete about their activity and depression levels.

Video: Qualitative Types and Experimental/Nonexperimental Research
In summary, there are two main approaches to research designs: Quantitative and qualitative. They each seek to answer questions, but quantitative research is meant to generalize its findings to the population whereas qualitative research seeks to understand phenomenon and develop theories about the human lived experiences.
References & Attribution
“ Light bulb doodle ” by rawpixel licensed CC0
“ Orange flame ” by rawpixel licensed CC0 .
Chen, P., Nunez-Smith, M., Bernheim, S… (2010). Professional experiences of international medical graduates practicing primary care in the United States. Journal of General Internal Medicine, 25 (9), 947-53.
Haedtke, C., Smith, M., VanBuren, J., Kein, D., Turvey, C. (2017). The relationships among pain, depression, and physical activity in patients with heart failure. Journal of Cardiovascular Nursing, 32 (5), E21-E25.
Pankong, O., Pothiban, L., Sucamvang, K., Khampolsiri, T. (2018). A randomized controlled trial of enhancing positive aspects of caregiving in Thai dementia caregivers for dementia. Pacific Rim Internal Journal of Nursing Res, 22 (2), 131-143.
Evidence-Based Practice & Research Methodologies Copyright © by Tracy Fawns is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License , except where otherwise noted.
Share This Book
Evidence-Based Medicine: Types of Studies
- What is Evidence-Based Practice?
- Question Types and Corresponding Resources
- Types of Studies
- Practice Guidelines
- Step 3: Appraise This link opens in a new window
- Steps 4-5: Apply & Assess
Experimental vs. Observational Studies
An observational study is a study in which the investigator cannot control the assignment of treatment to subjects because the participants or conditions are not directly assigned by the researcher.
- Examines predetermined treatments, interventions, policies, and their effects
- Four main types: case series , case-control studies , cross-sectional studies , and cohort studies
In an experimental study , the investigators directly manipulate or assign participants to different interventions or environments
Experimental studies that involve humans are called clinical trials . They fall into two categories: those with controls, and those without controls.
- Controlled trials - studies in which the experimental drug or procedure is compared with another drug or procedure
- Uncontrolled trials - studies in which the investigators' experience with the experimental drug or procedure is described, but the treatment is not compared with another treatment
Definitions taken from: Dawson B, Trapp R.G. (2004). Chapter 2. Study Designs in Medical Research. In Dawson B, Trapp R.G. (Eds), Basic & Clinical Biostatistics, 4e . Retrieved September 15, 2014 from https://accessmedicine.mhmedical.com/book.aspx?bookid=2724
Levels of Evidence Pyramid
Levels of Evidence Pyramid created by Andy Puro, September 2014
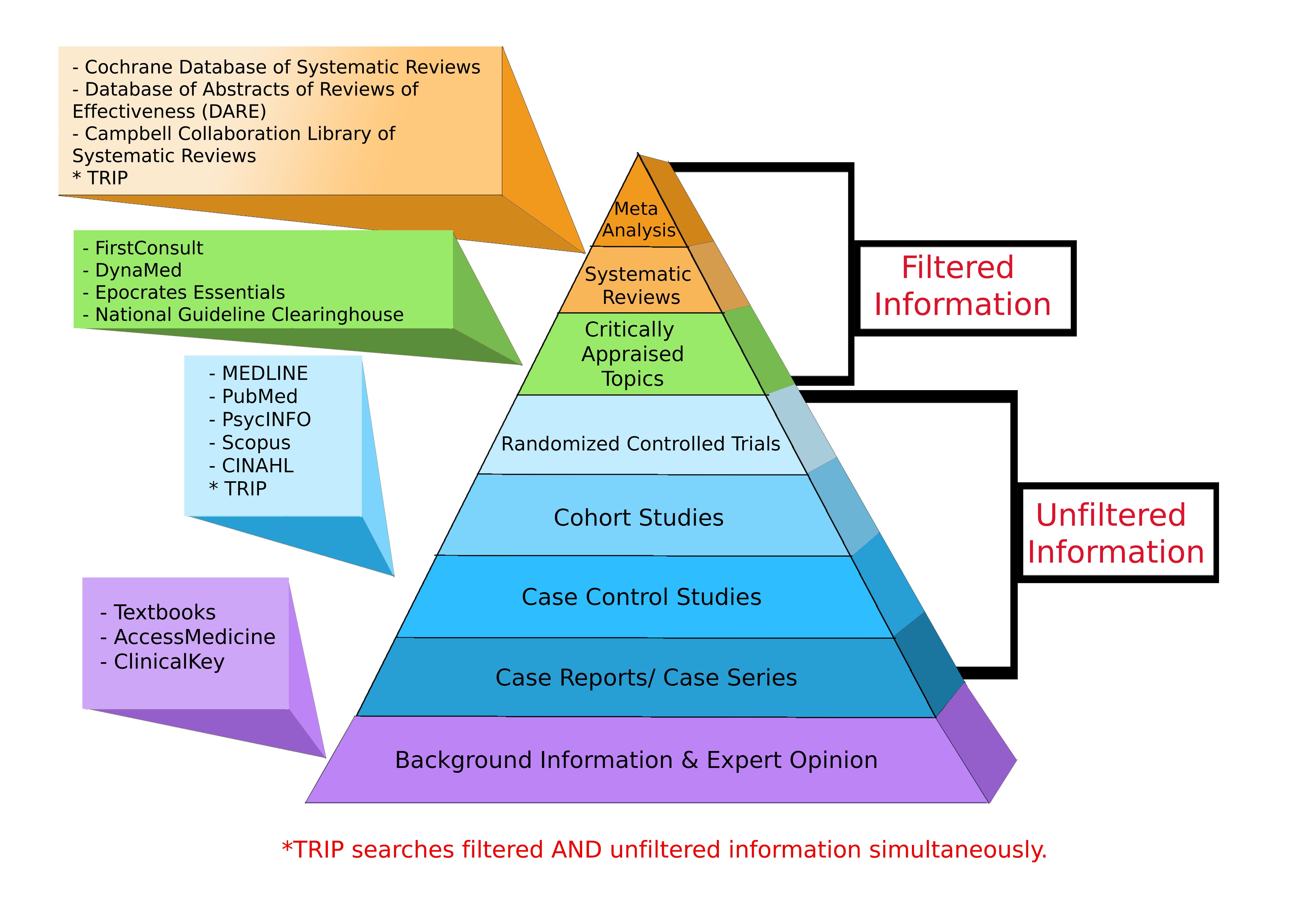
Additional Study Design Resources
Study Design 101 : Himmelfarb's tutorial on study types and how to find them
Study Designs (Centre for Evidence Based Medicine, University of Oxford)
Learn about Clinical Studies (ClinicalTrials.gov, National Institutes of Health)
Study Designs Guide (Deakin University)
- << Previous: Step 2: Acquire
- Next: Practice Guidelines >>

- Last Updated: Jul 1, 2024 9:31 AM
- URL: https://guides.himmelfarb.gwu.edu/ebm

- Himmelfarb Intranet
- Privacy Notice
- Terms of Use
- GW is committed to digital accessibility. If you experience a barrier that affects your ability to access content on this page, let us know via the Accessibility Feedback Form .
- Himmelfarb Health Sciences Library
- 2300 Eye St., NW, Washington, DC 20037
- Phone: (202) 994-2962
- [email protected]
- https://himmelfarb.gwu.edu
- Skip to main content
- Skip to FDA Search
- Skip to in this section menu
- Skip to footer links

The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you're on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
U.S. Food and Drug Administration
- Search
- Menu
- For Patients
- Clinical Trials: What Patients Need to Know
What Are the Different Types of Clinical Research?
Different types of clinical research are used depending on what the researchers are studying. Below are descriptions of some different kinds of clinical research.
Treatment Research generally involves an intervention such as medication, psychotherapy, new devices, or new approaches to surgery or radiation therapy.
Prevention Research looks for better ways to prevent disorders from developing or returning. Different kinds of prevention research may study medicines, vitamins, vaccines, minerals, or lifestyle changes.
Diagnostic Research refers to the practice of looking for better ways to identify a particular disorder or condition.
Screening Research aims to find the best ways to detect certain disorders or health conditions.
Quality of Life Research explores ways to improve comfort and the quality of life for individuals with a chronic illness.
Genetic studies aim to improve the prediction of disorders by identifying and understanding how genes and illnesses may be related. Research in this area may explore ways in which a person’s genes make him or her more or less likely to develop a disorder. This may lead to development of tailor-made treatments based on a patient’s genetic make-up.
Epidemiological studies seek to identify the patterns, causes, and control of disorders in groups of people.
An important note: some clinical research is “outpatient,” meaning that participants do not stay overnight at the hospital. Some is “inpatient,” meaning that participants will need to stay for at least one night in the hospital or research center. Be sure to ask the researchers what their study requires.
Phases of clinical trials: when clinical research is used to evaluate medications and devices Clinical trials are a kind of clinical research designed to evaluate and test new interventions such as psychotherapy or medications. Clinical trials are often conducted in four phases. The trials at each phase have a different purpose and help scientists answer different questions.
Phase I trials Researchers test an experimental drug or treatment in a small group of people for the first time. The researchers evaluate the treatment’s safety, determine a safe dosage range, and identify side effects.
Phase II trials The experimental drug or treatment is given to a larger group of people to see if it is effective and to further evaluate its safety.
Phase III trials The experimental study drug or treatment is given to large groups of people. Researchers confirm its effectiveness, monitor side effects, compare it to commonly used treatments, and collect information that will allow the experimental drug or treatment to be used safely.
Phase IV trials Post-marketing studies, which are conducted after a treatment is approved for use by the FDA, provide additional information including the treatment or drug’s risks, benefits, and best use.
Examples of other kinds of clinical research Many people believe that all clinical research involves testing of new medications or devices. This is not true, however. Some studies do not involve testing medications and a person’s regular medications may not need to be changed. Healthy volunteers are also needed so that researchers can compare their results to results of people with the illness being studied. Some examples of other kinds of research include the following:
A long-term study that involves psychological tests or brain scans
A genetic study that involves blood tests but no changes in medication
A study of family history that involves talking to family members to learn about people’s medical needs and history.
Research Study Types
There are many different types of research studies, and each has distinct strengths and weaknesses. In general, randomized trials and cohort studies provide the best information when looking at the link between a certain factor (like diet) and a health outcome (like heart disease).
Laboratory and Animal Studies
These are studies done in laboratories on cells, tissue, or animals.
- Strengths: Laboratories provide strictly controlled conditions and are often the genesis of scientific ideas that go on to have a broad impact on human health. They can help understand the mechanisms of disease.
- Weaknesses: Laboratory and animal studies are only a starting point. Animals or cells are not a substitute for humans.
Cross-Sectional Surveys
These studies examine the incidence of a certain outcome (disease or other health characteristic) in a specific group of people at one point in time. Surveys are often sent to participants to gather data about the outcome of interest.
- Strengths: Inexpensive and easy to perform.
- Weaknesses: Can only establish an association in that one specific time period.
Case-Control Studies
These studies look at the characteristics of one group of people who already have a certain health outcome (the cases) and compare them with a similar group of people who do not have the outcome (the controls). An example may be looking at a group of people with heart disease and another group without heart disease who are similar in age, sex, and economic status, and comparing their intakes of fruits and vegetables to see if this exposure could be associated with heart disease risk.
- Strengths: Case-control studies can be done quickly and relatively cheaply.
- Weaknesses: Not ideal for studying diet because they gather information from the past, which can be difficult for most people to recall accurately. Furthermore, people with illnesses often recall past behaviors differently from those without illness. This opens such studies to potential inaccuracy and bias in the information they gather.
Cohort Studies
These are observational studies that follow large groups of people over a long period of time, years or even decades, to find associations of an exposure(s) with disease outcomes. Researchers regularly gather information from the people in the study on several variables (like meat intake, physical activity level, and weight). Once a specified amount of time has elapsed, the characteristics of people in the group are compared to test specific hypotheses (such as a link between high versus low intake of carotenoid-rich foods and glaucoma, or high versus low meat intake and prostate cancer).
- Strengths: Participants are not required to change their diets or lifestyle as may be with randomized controlled studies. Study sizes may be larger than other study types. They generally provide more reliable information than case-control studies because they don’t rely on information from the past. Cohort studies gather information from participants at the beginning and throughout the study, long before they may develop the disease being studied. As a group, many of these types of studies have provided valuable information about the link between lifestyle factors and disease.
- Weaknesses: A longer duration of following participants make these studies time-consuming and expensive. Results cannot suggest cause-and-effect, only associations. Evaluation of dietary intake is self-reported.
Two of the largest and longest-running cohort studies of diet are the Harvard-based Nurses’ Health Study and the Health Professionals Follow-up Study.
If you follow nutrition news, chances are you have come across findings from a cohort called the Nurses’ Health Study . The Nurses’ Health Study (NHS) began in 1976, spearheaded by researchers from the Channing Laboratory at the Brigham and Women’s Hospital, Harvard Medical School, and the Harvard T.H. Chan School of Public Health, with funding from the National Institutes of Health. It gathered registered nurses ages 30-55 years from across the U.S. to respond to a series of questionnaires. Nurses were specifically chosen because of their ability to complete the health-related, often very technical, questionnaires thoroughly and accurately. They showed motivation to participate in the long-term study that required ongoing questionnaires every two years. Furthermore, the group provided blood, urine, and other samples over the course of the study.
The NHS is a prospective cohort study, meaning a group of people who are followed forward in time to examine lifestyle habits or other characteristics to see if they develop a disease, death, or some other indicated outcome. In comparison, a retrospective cohort study would specify a disease or outcome and look back in time at the group to see if there were common factors leading to the disease or outcome. A benefit of prospective studies over retrospective studies is greater accuracy in reporting details, such as food intake, that is not distorted by the diagnosis of illness.
To date, there are three NHS cohorts: NHS original cohort, NHS II, and NHS 3. Below are some features unique to each cohort.
NHS – Original Cohort
- Started in 1976 by Frank Speizer, M.D.
- Participants: 121,700 married women, ages 30 to 55 in 1976.
- Outcomes studied: Impact of contraceptive methods and smoking on breast cancer; later this was expanded to observe other lifestyle factors and behaviors in relation to 30 diseases.
- A food frequency questionnaire was added in 1980 to collect information on dietary intake, and continues to be collected every four years.
- Started in 1989 by Walter Willett, M.D., M.P.H., Dr.P.H., and colleagues.
- Participants: 116,430 single and married women, ages 25 to 42 in 1989.
- Outcomes studied: Impact on women’s health of oral contraceptives initiated during adolescence, diet and physical activity in adolescence, and lifestyle risk factors in a younger population than the NHS Original Cohort. The wide range of diseases examined in the original NHS is now also being studied in NHSII.
- The first food frequency questionnaire was collected in 1991, and is collected every four years.
- Started in 2010 by Jorge Chavarro, M.D., Sc.M., Sc.D, Walter Willett, M.D., M.P.H., Dr.P.H., Janet Rich-Edwards, Sc.D., M.P.H, and Stacey Missmer, Sc.D.
- Participants: Expanded to include not just registered nurses but licensed practical nurses (LPN) and licensed vocational nurses (LVN), ages 19 to 46. Enrollment is currently open.
- Inclusion of more diverse population of nurses, including male nurses and nurses from Canada.
- Outcomes studied: Dietary patterns, lifestyle, environment, and nursing occupational exposures that may impact men’s and women’s health; the impact of new hormone preparations and fertility/pregnancy on women’s health; relationship of diet in adolescence on breast cancer risk.
From these three cohorts, extensive research has been published regarding the association of diet, smoking, physical activity levels, overweight and obesity, oral contraceptive use, hormone therapy, endogenous hormones, dietary factors, sleep, genetics, and other behaviors and characteristics with various diseases. In 2016, in celebration of the 40 th Anniversary of NHS, the American Journal of Public Health’s September issue was dedicated to featuring the many contributions of the Nurses’ Health Studies to public health.
Growing Up Today Study (GUTS)
In 1996, recruitment began for a new cross-generational cohort called GUTS (Growing Up Today Study) —children of nurses from the NHS II. GUTS is composed of 27,802 girls and boys who were between the ages of 9 and 17 at the time of enrollment. As the entire cohort has entered adulthood, they complete annual questionnaires including information on dietary intake, weight changes, exercise level, substance and alcohol use, body image, and environmental factors. Researchers are looking at conditions more common in young adults such as asthma, skin cancer, eating disorders, and sports injuries.
Randomized Trials
Like cohort studies, these studies follow a group of people over time. However, with randomized trials, the researchers intervene with a specific behavior change or treatment (such as following a specific diet or taking a supplement) to see how it affects a health outcome. They are called “randomized trials” because people in the study are randomly assigned to either receive or not receive the intervention. This randomization helps researchers determine the true effect the intervention has on the health outcome. Those who do not receive the intervention or labelled the “control group,” which means these participants do not change their behavior, or if the study is examining the effects of a vitamin supplement, the control group participants receive a placebo supplement that contains no active ingredients.
- Strengths: Considered the “gold standard” and best for determining the effectiveness of an intervention (e.g., dietary pattern, supplement) on an endpoint such as cancer or heart disease. Conducted in a highly controlled setting with limited variables that could affect the outcome. They determine cause-and-effect relationships.
- Weaknesses: High cost, potentially low long-term compliance with prescribed diets, and possible ethical issues. Due to expense, the study size may be small.
Meta-Analyses and Systematic Reviews
A meta-analysis collects data from several previous studies on one topic to analyze and combine the results using statistical methods to provide a summary conclusion. Meta-analyses are usually conducted using randomized controlled trials and cohort studies that have higher quality of evidence than other designs. A systematic review also examines past literature related to a specific topic and design, analyzing the quality of studies and results but may not pool the data. Sometimes a systematic review is followed by conducting a meta-analysis if the quality of the studies is good and the data can be combined.
- Strengths: Inexpensive and provides a general comprehensive summary of existing research on a topic. This can create an explanation or assumption to be used for further investigation.
- Weaknesses: Prone to selection bias, as the authors can choose or exclude certain studies, which can change the resulting outcome. Combining data that includes lower-quality studies can also skew the results.
A primer on systematic review and meta-analysis in diabetes research
Terms of use.
The contents of this website are for educational purposes and are not intended to offer personal medical advice. You should seek the advice of your physician or other qualified health provider with any questions you may have regarding a medical condition. Never disregard professional medical advice or delay in seeking it because of something you have read on this website. The Nutrition Source does not recommend or endorse any products.
15 Types of Research Methods

Chris Drew (PhD)
Dr. Chris Drew is the founder of the Helpful Professor. He holds a PhD in education and has published over 20 articles in scholarly journals. He is the former editor of the Journal of Learning Development in Higher Education. [Image Descriptor: Photo of Chris]
Learn about our Editorial Process

Research methods refer to the strategies, tools, and techniques used to gather and analyze data in a structured way in order to answer a research question or investigate a hypothesis (Hammond & Wellington, 2020).
Generally, we place research methods into two categories: quantitative and qualitative. Each has its own strengths and weaknesses, which we can summarize as:
- Quantitative research can achieve generalizability through scrupulous statistical analysis applied to large sample sizes.
- Qualitative research achieves deep, detailed, and nuance accounts of specific case studies, which are not generalizable.
Some researchers, with the aim of making the most of both quantitative and qualitative research, employ mixed methods, whereby they will apply both types of research methods in the one study, such as by conducting a statistical survey alongside in-depth interviews to add context to the quantitative findings.
Below, I’ll outline 15 common research methods, and include pros, cons, and examples of each .
Types of Research Methods
Research methods can be broadly categorized into two types: quantitative and qualitative.
- Quantitative methods involve systematic empirical investigation of observable phenomena via statistical, mathematical, or computational techniques, providing an in-depth understanding of a specific concept or phenomenon (Schweigert, 2021). The strengths of this approach include its ability to produce reliable results that can be generalized to a larger population, although it can lack depth and detail.
- Qualitative methods encompass techniques that are designed to provide a deep understanding of a complex issue, often in a specific context, through collection of non-numerical data (Tracy, 2019). This approach often provides rich, detailed insights but can be time-consuming and its findings may not be generalizable.
These can be further broken down into a range of specific research methods and designs:
| Primarily Quantitative Methods | Primarily Qualitative methods |
|---|---|
| Experimental Research | Case Study |
| Surveys and Questionnaires | Ethnography |
| Longitudinal Studies | Phenomenology |
| Cross-Sectional Studies | Historical research |
| Correlational Research | Content analysis |
| Causal-Comparative Research | Grounded theory |
| Meta-Analysis | Action research |
| Quasi-Experimental Design | Observational research |
Combining the two methods above, mixed methods research mixes elements of both qualitative and quantitative research methods, providing a comprehensive understanding of the research problem . We can further break these down into:
- Sequential Explanatory Design (QUAN→QUAL): This methodology involves conducting quantitative analysis first, then supplementing it with a qualitative study.
- Sequential Exploratory Design (QUAL→QUAN): This methodology goes in the other direction, starting with qualitative analysis and ending with quantitative analysis.
Let’s explore some methods and designs from both quantitative and qualitative traditions, starting with qualitative research methods.
Qualitative Research Methods
Qualitative research methods allow for the exploration of phenomena in their natural settings, providing detailed, descriptive responses and insights into individuals’ experiences and perceptions (Howitt, 2019).
These methods are useful when a detailed understanding of a phenomenon is sought.
1. Ethnographic Research
Ethnographic research emerged out of anthropological research, where anthropologists would enter into a setting for a sustained period of time, getting to know a cultural group and taking detailed observations.
Ethnographers would sometimes even act as participants in the group or culture, which many scholars argue is a weakness because it is a step away from achieving objectivity (Stokes & Wall, 2017).
In fact, at its most extreme version, ethnographers even conduct research on themselves, in a fascinating methodology call autoethnography .
The purpose is to understand the culture, social structure, and the behaviors of the group under study. It is often useful when researchers seek to understand shared cultural meanings and practices in their natural settings.
However, it can be time-consuming and may reflect researcher biases due to the immersion approach.
| Pros of Ethnographic Research | Cons of Ethnographic Research |
|---|---|
| 1. Provides deep cultural insights | 1. Time-consuming |
| 2. Contextually relevant findings | 2. Potential researcher bias |
| 3. Explores dynamic social processes | 3. May |
Example of Ethnography
Liquidated: An Ethnography of Wall Street by Karen Ho involves an anthropologist who embeds herself with Wall Street firms to study the culture of Wall Street bankers and how this culture affects the broader economy and world.
2. Phenomenological Research
Phenomenological research is a qualitative method focused on the study of individual experiences from the participant’s perspective (Tracy, 2019).
It focuses specifically on people’s experiences in relation to a specific social phenomenon ( see here for examples of social phenomena ).
This method is valuable when the goal is to understand how individuals perceive, experience, and make meaning of particular phenomena. However, because it is subjective and dependent on participants’ self-reports, findings may not be generalizable, and are highly reliant on self-reported ‘thoughts and feelings’.
| Pros of Phenomenological Research | Cons of Phenomenological Research |
|---|---|
| 1. Provides rich, detailed data | 1. Limited generalizability |
| 2. Highlights personal experience and perceptions | 2. Data collection can be time-consuming |
| 3. Allows exploration of complex phenomena | 3. Requires highly skilled researchers |
Example of Phenomenological Research
A phenomenological approach to experiences with technology by Sebnem Cilesiz represents a good starting-point for formulating a phenomenological study. With its focus on the ‘essence of experience’, this piece presents methodological, reliability, validity, and data analysis techniques that phenomenologists use to explain how people experience technology in their everyday lives.
3. Historical Research
Historical research is a qualitative method involving the examination of past events to draw conclusions about the present or make predictions about the future (Stokes & Wall, 2017).
As you might expect, it’s common in the research branches of history departments in universities.
This approach is useful in studies that seek to understand the past to interpret present events or trends. However, it relies heavily on the availability and reliability of source materials, which may be limited.
Common data sources include cultural artifacts from both material and non-material culture , which are then examined, compared, contrasted, and contextualized to test hypotheses and generate theories.
| Pros of Historical Research | Cons of Historical Research |
|---|---|
| 1. | 1. Dependent on available sources |
| 2. Can help understand current events or trends | 2. Potential bias in source materials |
| 3. Allows the study of change over time | 3. Difficult to replicate |
Example of Historical Research
A historical research example might be a study examining the evolution of gender roles over the last century. This research might involve the analysis of historical newspapers, advertisements, letters, and company documents, as well as sociocultural contexts.
4. Content Analysis
Content analysis is a research method that involves systematic and objective coding and interpreting of text or media to identify patterns, themes, ideologies, or biases (Schweigert, 2021).
A content analysis is useful in analyzing communication patterns, helping to reveal how texts such as newspapers, movies, films, political speeches, and other types of ‘content’ contain narratives and biases.
However, interpretations can be very subjective, which often requires scholars to engage in practices such as cross-comparing their coding with peers or external researchers.
Content analysis can be further broken down in to other specific methodologies such as semiotic analysis, multimodal analysis , and discourse analysis .
| Pros of Content Analysis | Cons of Content Analysis |
|---|---|
| 1. Unobtrusive data collection | 1. Lacks contextual information |
| 2. Allows for large sample analysis | 2. Potential coder bias |
| 3. Replicable and reliable if done properly | 3. May overlook nuances |
Example of Content Analysis
How is Islam Portrayed in Western Media? by Poorebrahim and Zarei (2013) employs a type of content analysis called critical discourse analysis (common in poststructuralist and critical theory research ). This study by Poorebrahum and Zarei combs through a corpus of western media texts to explore the language forms that are used in relation to Islam and Muslims, finding that they are overly stereotyped, which may represent anti-Islam bias or failure to understand the Islamic world.
5. Grounded Theory Research
Grounded theory involves developing a theory during and after data collection rather than beforehand.
This is in contrast to most academic research studies, which start with a hypothesis or theory and then testing of it through a study, where we might have a null hypothesis (disproving the theory) and an alternative hypothesis (supporting the theory).
Grounded Theory is useful because it keeps an open mind to what the data might reveal out of the research. It can be time-consuming and requires rigorous data analysis (Tracy, 2019).
| Pros of Grounded Theory Research | Cons of Grounded Theory Research |
|---|---|
| 1. Helps with theory development | 1. Time-consuming |
| 2. Rigorous data analysis | 2. Requires iterative data collection and analysis |
| 3. Can fill gaps in existing theories | 3. Requires skilled researchers |
Grounded Theory Example
Developing a Leadership Identity by Komives et al (2005) employs a grounded theory approach to develop a thesis based on the data rather than testing a hypothesis. The researchers studied the leadership identity of 13 college students taking on leadership roles. Based on their interviews, the researchers theorized that the students’ leadership identities shifted from a hierarchical view of leadership to one that embraced leadership as a collaborative concept.
6. Action Research
Action research is an approach which aims to solve real-world problems and bring about change within a setting. The study is designed to solve a specific problem – or in other words, to take action (Patten, 2017).
This approach can involve mixed methods, but is generally qualitative because it usually involves the study of a specific case study wherein the researcher works, e.g. a teacher studying their own classroom practice to seek ways they can improve.
Action research is very common in fields like education and nursing where practitioners identify areas for improvement then implement a study in order to find paths forward.
| Pros of Action Research | Cons of Action Research |
|---|---|
| 1. Addresses real-world problems and seeks to find solutions. | 1. It is time-consuming and often hard to implement into a practitioner’s already busy schedule |
| 2. Integrates research and action in an action-research cycle. | 2. Requires collaboration between researcher, practitioner, and research participants. |
| 3. Can bring about positive change in isolated instances, such as in a school or nursery setting. | 3. Complexity of managing dual roles (where the researcher is also often the practitioner) |
Action Research Example
Using Digital Sandbox Gaming to Improve Creativity Within Boys’ Writing by Ellison and Drew was a research study one of my research students completed in his own classroom under my supervision. He implemented a digital game-based approach to literacy teaching with boys and interviewed his students to see if the use of games as stimuli for storytelling helped draw them into the learning experience.
7. Natural Observational Research
Observational research can also be quantitative (see: experimental research), but in naturalistic settings for the social sciences, researchers tend to employ qualitative data collection methods like interviews and field notes to observe people in their day-to-day environments.
This approach involves the observation and detailed recording of behaviors in their natural settings (Howitt, 2019). It can provide rich, in-depth information, but the researcher’s presence might influence behavior.
While observational research has some overlaps with ethnography (especially in regard to data collection techniques), it tends not to be as sustained as ethnography, e.g. a researcher might do 5 observations, every second Monday, as opposed to being embedded in an environment.
| Pros of Qualitative Observational Research | Cons of Qualitative Observational Research |
|---|---|
| 1. Captures behavior in natural settings, allowing for interesting insights into authentic behaviors. | 1. Researcher’s presence may influence behavior |
| 2. Can provide rich, detailed data through the researcher’s vignettes. | 2. Can be time-consuming |
| 3. Non-invasive because researchers want to observe natural activities rather than interfering with research participants. | 3. Requires skilled and trained observers |
Observational Research Example
A researcher might use qualitative observational research to study the behaviors and interactions of children at a playground. The researcher would document the behaviors observed, such as the types of games played, levels of cooperation , and instances of conflict.
8. Case Study Research
Case study research is a qualitative method that involves a deep and thorough investigation of a single individual, group, or event in order to explore facets of that phenomenon that cannot be captured using other methods (Stokes & Wall, 2017).
Case study research is especially valuable in providing contextualized insights into specific issues, facilitating the application of abstract theories to real-world situations (Patten, 2017).
However, findings from a case study may not be generalizable due to the specific context and the limited number of cases studied (Walliman, 2021).
| Pros of Case Study Research | Cons of Case Study Research |
|---|---|
| 1. Provides detailed insights | 1. Limited generalizability |
| 2. Facilitates the study of complex phenomena | 2. Can be time-consuming |
| 3. Can test or generate theories | 3. Subject to observer bias |
See More: Case Study Advantages and Disadvantages
Example of a Case Study
Scholars conduct a detailed exploration of the implementation of a new teaching method within a classroom setting. The study focuses on how the teacher and students adapt to the new method, the challenges encountered, and the outcomes on student performance and engagement. While the study provides specific and detailed insights of the teaching method in that classroom, it cannot be generalized to other classrooms, as statistical significance has not been established through this qualitative approach.
Quantitative Research Methods
Quantitative research methods involve the systematic empirical investigation of observable phenomena via statistical, mathematical, or computational techniques (Pajo, 2022). The focus is on gathering numerical data and generalizing it across groups of people or to explain a particular phenomenon.
9. Experimental Research
Experimental research is a quantitative method where researchers manipulate one variable to determine its effect on another (Walliman, 2021).
This is common, for example, in high-school science labs, where students are asked to introduce a variable into a setting in order to examine its effect.
This type of research is useful in situations where researchers want to determine causal relationships between variables. However, experimental conditions may not reflect real-world conditions.
| Pros of Experimental Research | Cons of Experimental Research |
|---|---|
| 1. Allows for determination of causality | 1. Might not reflect real-world conditions |
| 2. Allows for the study of phenomena in highly controlled environments to minimize research contamination. | 2. Can be costly and time-consuming to create a controlled environment. |
| 3. Can be replicated so other researchers can test and verify the results. | 3. Ethical concerns need to be addressed as the research is directly manipulating variables. |
Example of Experimental Research
A researcher may conduct an experiment to determine the effects of a new educational approach on student learning outcomes. Students would be randomly assigned to either the control group (traditional teaching method) or the experimental group (new educational approach).
10. Surveys and Questionnaires
Surveys and questionnaires are quantitative methods that involve asking research participants structured and predefined questions to collect data about their attitudes, beliefs, behaviors, or characteristics (Patten, 2017).
Surveys are beneficial for collecting data from large samples, but they depend heavily on the honesty and accuracy of respondents.
They tend to be seen as more authoritative than their qualitative counterparts, semi-structured interviews, because the data is quantifiable (e.g. a questionnaire where information is presented on a scale from 1 to 10 can allow researchers to determine and compare statistical means, averages, and variations across sub-populations in the study).
| Pros of Surveys and Questionnaires | Cons of Surveys and Questionnaires |
|---|---|
| 1. Data can be gathered from larger samples than is possible in qualitative research. | 1. There is heavy dependence on respondent honesty |
| 2. The data is quantifiable, allowing for comparison across subpopulations | 2. There is limited depth of response as opposed to qualitative approaches. |
| 3. Can be cost-effective and time-efficient | 3. Static with no flexibility to explore responses (unlike semi- or unstrcutured interviewing) |
Example of a Survey Study
A company might use a survey to gather data about employee job satisfaction across its offices worldwide. Employees would be asked to rate various aspects of their job satisfaction on a Likert scale. While this method provides a broad overview, it may lack the depth of understanding possible with other methods (Stokes & Wall, 2017).
11. Longitudinal Studies
Longitudinal studies involve repeated observations of the same variables over extended periods (Howitt, 2019). These studies are valuable for tracking development and change but can be costly and time-consuming.
With multiple data points collected over extended periods, it’s possible to examine continuous changes within things like population dynamics or consumer behavior. This makes a detailed analysis of change possible.
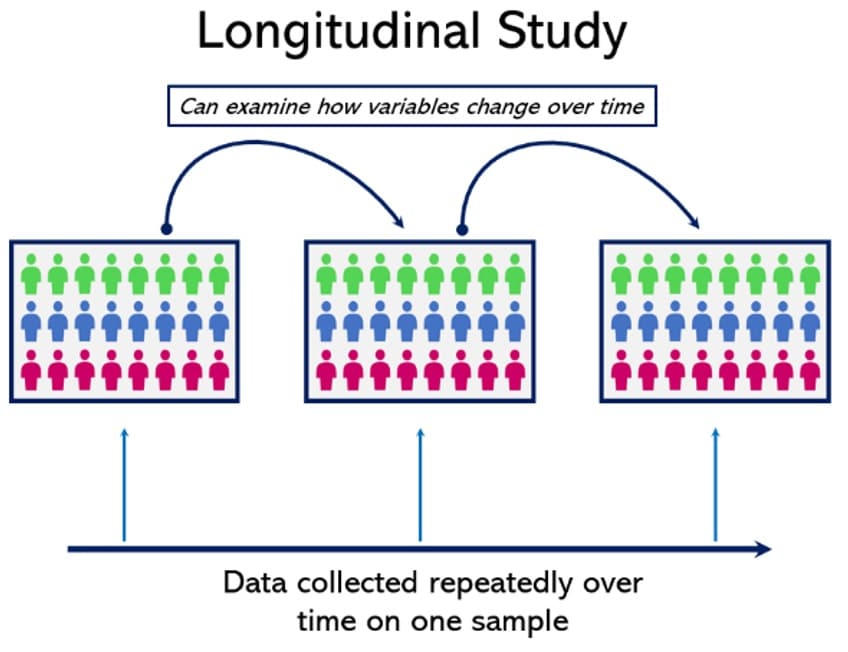
Perhaps the most relatable example of a longitudinal study is a national census, which is taken on the same day every few years, to gather comparative demographic data that can show how a nation is changing over time.
While longitudinal studies are commonly quantitative, there are also instances of qualitative ones as well, such as the famous 7 Up study from the UK, which studies 14 individuals every 7 years to explore their development over their lives.
| Pros of Longitudinal Studies | Cons of Longitudinal Studies |
|---|---|
| 1. Tracks changes over time allowing for comparison of past to present events. | 1. Is almost by definition time-consuming because time needs to pass between each data collection session. |
| 2. Can identify sequences of events, but causality is often harder to determine. | 2. There is high risk of participant dropout over time as participants move on with their lives. |
Example of a Longitudinal Study
A national census, taken every few years, uses surveys to develop longitudinal data, which is then compared and analyzed to present accurate trends over time. Trends a census can reveal include changes in religiosity, values and attitudes on social issues, and much more.
12. Cross-Sectional Studies
Cross-sectional studies are a quantitative research method that involves analyzing data from a population at a specific point in time (Patten, 2017). They provide a snapshot of a situation but cannot determine causality.
This design is used to measure and compare the prevalence of certain characteristics or outcomes in different groups within the sampled population.
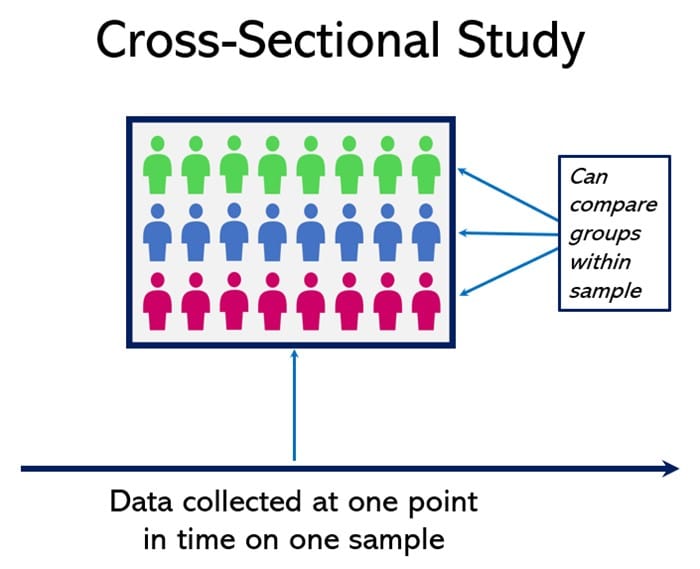
The major advantage of cross-sectional design is its ability to measure a wide range of variables simultaneously without needing to follow up with participants over time.
However, cross-sectional studies do have limitations . This design can only show if there are associations or correlations between different variables, but cannot prove cause and effect relationships, temporal sequence, changes, and trends over time.
| Pros of Cross-Sectional Studies | Cons of Cross-Sectional Studies |
|---|---|
| 1. Quick and inexpensive, with no long-term commitment required. | 1. Cannot determine causality because it is a simple snapshot, with no time delay between data collection points. |
| 2. Good for descriptive analyses. | 2. Does not allow researchers to follow up with research participants. |
Example of a Cross-Sectional Study
Our longitudinal study example of a national census also happens to contain cross-sectional design. One census is cross-sectional, displaying only data from one point in time. But when a census is taken once every few years, it becomes longitudinal, and so long as the data collection technique remains unchanged, identification of changes will be achievable, adding another time dimension on top of a basic cross-sectional study.
13. Correlational Research
Correlational research is a quantitative method that seeks to determine if and to what degree a relationship exists between two or more quantifiable variables (Schweigert, 2021).
This approach provides a fast and easy way to make initial hypotheses based on either positive or negative correlation trends that can be observed within dataset.
While correlational research can reveal relationships between variables, it cannot establish causality.
Methods used for data analysis may include statistical correlations such as Pearson’s or Spearman’s.
| Pros of Correlational Research | Cons of Correlational Research |
|---|---|
| 1. Reveals relationships between variables | 1. Cannot determine causality |
| 2. Can use existing data | 2. May be |
| 3. Can guide further experimental research | 3. Correlation may be coincidental |
Example of Correlational Research
A team of researchers is interested in studying the relationship between the amount of time students spend studying and their academic performance. They gather data from a high school, measuring the number of hours each student studies per week and their grade point averages (GPAs) at the end of the semester. Upon analyzing the data, they find a positive correlation, suggesting that students who spend more time studying tend to have higher GPAs.
14. Quasi-Experimental Design Research
Quasi-experimental design research is a quantitative research method that is similar to experimental design but lacks the element of random assignment to treatment or control.
Instead, quasi-experimental designs typically rely on certain other methods to control for extraneous variables.
The term ‘quasi-experimental’ implies that the experiment resembles a true experiment, but it is not exactly the same because it doesn’t meet all the criteria for a ‘true’ experiment, specifically in terms of control and random assignment.
Quasi-experimental design is useful when researchers want to study a causal hypothesis or relationship, but practical or ethical considerations prevent them from manipulating variables and randomly assigning participants to conditions.
| Pros | Cons |
|---|---|
| 1. It’s more feasible to implement than true experiments. | 1. Without random assignment, it’s harder to rule out confounding variables. |
| 2. It can be conducted in real-world settings, making the findings more applicable to the real world. | 2. The lack of random assignment may of the study. |
| 3. Useful when it’s unethical or impossible to manipulate the independent variable or randomly assign participants. | 3. It’s more difficult to establish a cause-effect relationship due to the potential for confounding variables. |
Example of Quasi-Experimental Design
A researcher wants to study the impact of a new math tutoring program on student performance. However, ethical and practical constraints prevent random assignment to the “tutoring” and “no tutoring” groups. Instead, the researcher compares students who chose to receive tutoring (experimental group) to similar students who did not choose to receive tutoring (control group), controlling for other variables like grade level and previous math performance.
Related: Examples and Types of Random Assignment in Research
15. Meta-Analysis Research
Meta-analysis statistically combines the results of multiple studies on a specific topic to yield a more precise estimate of the effect size. It’s the gold standard of secondary research .
Meta-analysis is particularly useful when there are numerous studies on a topic, and there is a need to integrate the findings to draw more reliable conclusions.
Some meta-analyses can identify flaws or gaps in a corpus of research, when can be highly influential in academic research, despite lack of primary data collection.
However, they tend only to be feasible when there is a sizable corpus of high-quality and reliable studies into a phenomenon.
| Pros | Cons |
|---|---|
| Increased Statistical Power: By combining data from multiple studies, meta-analysis increases the statistical power to detect effects. | Publication Bias: Studies with null or negative findings are less likely to be published, leading to an overestimation of effect sizes. |
| Greater Precision: It provides more precise estimates of effect sizes by reducing the influence of random error. | Quality of Studies: of a meta-analysis depends on the quality of the studies included. |
| Resolving Discrepancies: Meta-analysis can help resolve disagreements between different studies on a topic. | Heterogeneity: Differences in study design, sample, or procedures can introduce heterogeneity, complicating interpretation of results. |
Example of a Meta-Analysis
The power of feedback revisited (Wisniewski, Zierer & Hattie, 2020) is a meta-analysis that examines 435 empirical studies research on the effects of feedback on student learning. They use a random-effects model to ascertain whether there is a clear effect size across the literature. The authors find that feedback tends to impact cognitive and motor skill outcomes but has less of an effect on motivational and behavioral outcomes.
Choosing a research method requires a lot of consideration regarding what you want to achieve, your research paradigm, and the methodology that is most valuable for what you are studying. There are multiple types of research methods, many of which I haven’t been able to present here. Generally, it’s recommended that you work with an experienced researcher or research supervisor to identify a suitable research method for your study at hand.
Hammond, M., & Wellington, J. (2020). Research methods: The key concepts . New York: Routledge.
Howitt, D. (2019). Introduction to qualitative research methods in psychology . London: Pearson UK.
Pajo, B. (2022). Introduction to research methods: A hands-on approach . New York: Sage Publications.
Patten, M. L. (2017). Understanding research methods: An overview of the essentials . New York: Sage
Schweigert, W. A. (2021). Research methods in psychology: A handbook . Los Angeles: Waveland Press.
Stokes, P., & Wall, T. (2017). Research methods . New York: Bloomsbury Publishing.
Tracy, S. J. (2019). Qualitative research methods: Collecting evidence, crafting analysis, communicating impact . London: John Wiley & Sons.
Walliman, N. (2021). Research methods: The basics. London: Routledge.

- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd-2/ 10 Reasons you’re Perpetually Single
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd-2/ 20 Montessori Toddler Bedrooms (Design Inspiration)
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd-2/ 21 Montessori Homeschool Setups
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd-2/ 101 Hidden Talents Examples
Leave a Comment Cancel Reply
Your email address will not be published. Required fields are marked *

Please consult the following sources for more information on these types of studies and terminology related to the studies.
- APA Style JARS Supplemental Glossary This webpage provides supplemental information on the terms used in APA Style JARS. This glossary is meant to supplement Chapter 3 of the Publication Manual of the American Psychological Association, Seventh Edition. It is not an exhaustive list of all terms employed in quantitative, qualitative, or mixed methods research, nor does it include all possible definitions for each term; definitions in addition to or different from those reported in this glossary may be found in other sources.
- APA Dictionary of Psychology More than 25,000 authoritative entries across 90 subfields of psychology.
- << Previous: Types of Research/Scholarly Articles
- Next: Research Methods & Analysis Resources >>
- Last Updated: Aug 19, 2024 2:23 PM
- URL: https://libguides.ggc.edu/Research_Statistics_Writing_Dissertation_Publishing
Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, generate accurate citations for free.
- Knowledge Base
Methodology

Research Methods | Definitions, Types, Examples
Research methods are specific procedures for collecting and analyzing data. Developing your research methods is an integral part of your research design . When planning your methods, there are two key decisions you will make.
First, decide how you will collect data . Your methods depend on what type of data you need to answer your research question :
- Qualitative vs. quantitative : Will your data take the form of words or numbers?
- Primary vs. secondary : Will you collect original data yourself, or will you use data that has already been collected by someone else?
- Descriptive vs. experimental : Will you take measurements of something as it is, or will you perform an experiment?
Second, decide how you will analyze the data .
- For quantitative data, you can use statistical analysis methods to test relationships between variables.
- For qualitative data, you can use methods such as thematic analysis to interpret patterns and meanings in the data.
Table of contents
Methods for collecting data, examples of data collection methods, methods for analyzing data, examples of data analysis methods, other interesting articles, frequently asked questions about research methods.
Data is the information that you collect for the purposes of answering your research question . The type of data you need depends on the aims of your research.
Qualitative vs. quantitative data
Your choice of qualitative or quantitative data collection depends on the type of knowledge you want to develop.
For questions about ideas, experiences and meanings, or to study something that can’t be described numerically, collect qualitative data .
If you want to develop a more mechanistic understanding of a topic, or your research involves hypothesis testing , collect quantitative data .
| Qualitative | to broader populations. . | |
|---|---|---|
| Quantitative | . |
You can also take a mixed methods approach , where you use both qualitative and quantitative research methods.
Primary vs. secondary research
Primary research is any original data that you collect yourself for the purposes of answering your research question (e.g. through surveys , observations and experiments ). Secondary research is data that has already been collected by other researchers (e.g. in a government census or previous scientific studies).
If you are exploring a novel research question, you’ll probably need to collect primary data . But if you want to synthesize existing knowledge, analyze historical trends, or identify patterns on a large scale, secondary data might be a better choice.
| Primary | . | methods. |
|---|---|---|
| Secondary |
Descriptive vs. experimental data
In descriptive research , you collect data about your study subject without intervening. The validity of your research will depend on your sampling method .
In experimental research , you systematically intervene in a process and measure the outcome. The validity of your research will depend on your experimental design .
To conduct an experiment, you need to be able to vary your independent variable , precisely measure your dependent variable, and control for confounding variables . If it’s practically and ethically possible, this method is the best choice for answering questions about cause and effect.
| Descriptive | . . | |
|---|---|---|
| Experimental |
Prevent plagiarism. Run a free check.
| Research method | Primary or secondary? | Qualitative or quantitative? | When to use |
|---|---|---|---|
| Primary | Quantitative | To test cause-and-effect relationships. | |
| Primary | Quantitative | To understand general characteristics of a population. | |
| Interview/focus group | Primary | Qualitative | To gain more in-depth understanding of a topic. |
| Observation | Primary | Either | To understand how something occurs in its natural setting. |
| Secondary | Either | To situate your research in an existing body of work, or to evaluate trends within a research topic. | |
| Either | Either | To gain an in-depth understanding of a specific group or context, or when you don’t have the resources for a large study. |
Your data analysis methods will depend on the type of data you collect and how you prepare it for analysis.
Data can often be analyzed both quantitatively and qualitatively. For example, survey responses could be analyzed qualitatively by studying the meanings of responses or quantitatively by studying the frequencies of responses.
Qualitative analysis methods
Qualitative analysis is used to understand words, ideas, and experiences. You can use it to interpret data that was collected:
- From open-ended surveys and interviews , literature reviews , case studies , ethnographies , and other sources that use text rather than numbers.
- Using non-probability sampling methods .
Qualitative analysis tends to be quite flexible and relies on the researcher’s judgement, so you have to reflect carefully on your choices and assumptions and be careful to avoid research bias .
Quantitative analysis methods
Quantitative analysis uses numbers and statistics to understand frequencies, averages and correlations (in descriptive studies) or cause-and-effect relationships (in experiments).
You can use quantitative analysis to interpret data that was collected either:
- During an experiment .
- Using probability sampling methods .
Because the data is collected and analyzed in a statistically valid way, the results of quantitative analysis can be easily standardized and shared among researchers.
| Research method | Qualitative or quantitative? | When to use |
|---|---|---|
| Quantitative | To analyze data collected in a statistically valid manner (e.g. from experiments, surveys, and observations). | |
| Meta-analysis | Quantitative | To statistically analyze the results of a large collection of studies. Can only be applied to studies that collected data in a statistically valid manner. |
| Qualitative | To analyze data collected from interviews, , or textual sources. To understand general themes in the data and how they are communicated. | |
| Either | To analyze large volumes of textual or visual data collected from surveys, literature reviews, or other sources. Can be quantitative (i.e. frequencies of words) or qualitative (i.e. meanings of words). |
Here's why students love Scribbr's proofreading services
Discover proofreading & editing
If you want to know more about statistics , methodology , or research bias , make sure to check out some of our other articles with explanations and examples.
- Chi square test of independence
- Statistical power
- Descriptive statistics
- Degrees of freedom
- Pearson correlation
- Null hypothesis
- Double-blind study
- Case-control study
- Research ethics
- Data collection
- Hypothesis testing
- Structured interviews
Research bias
- Hawthorne effect
- Unconscious bias
- Recall bias
- Halo effect
- Self-serving bias
- Information bias
Quantitative research deals with numbers and statistics, while qualitative research deals with words and meanings.
Quantitative methods allow you to systematically measure variables and test hypotheses . Qualitative methods allow you to explore concepts and experiences in more detail.
In mixed methods research , you use both qualitative and quantitative data collection and analysis methods to answer your research question .
A sample is a subset of individuals from a larger population . Sampling means selecting the group that you will actually collect data from in your research. For example, if you are researching the opinions of students in your university, you could survey a sample of 100 students.
In statistics, sampling allows you to test a hypothesis about the characteristics of a population.
The research methods you use depend on the type of data you need to answer your research question .
- If you want to measure something or test a hypothesis , use quantitative methods . If you want to explore ideas, thoughts and meanings, use qualitative methods .
- If you want to analyze a large amount of readily-available data, use secondary data. If you want data specific to your purposes with control over how it is generated, collect primary data.
- If you want to establish cause-and-effect relationships between variables , use experimental methods. If you want to understand the characteristics of a research subject, use descriptive methods.
Methodology refers to the overarching strategy and rationale of your research project . It involves studying the methods used in your field and the theories or principles behind them, in order to develop an approach that matches your objectives.
Methods are the specific tools and procedures you use to collect and analyze data (for example, experiments, surveys , and statistical tests ).
In shorter scientific papers, where the aim is to report the findings of a specific study, you might simply describe what you did in a methods section .
In a longer or more complex research project, such as a thesis or dissertation , you will probably include a methodology section , where you explain your approach to answering the research questions and cite relevant sources to support your choice of methods.
Is this article helpful?
Other students also liked, writing strong research questions | criteria & examples.
- What Is a Research Design | Types, Guide & Examples
- Data Collection | Definition, Methods & Examples
More interesting articles
- Between-Subjects Design | Examples, Pros, & Cons
- Cluster Sampling | A Simple Step-by-Step Guide with Examples
- Confounding Variables | Definition, Examples & Controls
- Construct Validity | Definition, Types, & Examples
- Content Analysis | Guide, Methods & Examples
- Control Groups and Treatment Groups | Uses & Examples
- Control Variables | What Are They & Why Do They Matter?
- Correlation vs. Causation | Difference, Designs & Examples
- Correlational Research | When & How to Use
- Critical Discourse Analysis | Definition, Guide & Examples
- Cross-Sectional Study | Definition, Uses & Examples
- Descriptive Research | Definition, Types, Methods & Examples
- Ethical Considerations in Research | Types & Examples
- Explanatory and Response Variables | Definitions & Examples
- Explanatory Research | Definition, Guide, & Examples
- Exploratory Research | Definition, Guide, & Examples
- External Validity | Definition, Types, Threats & Examples
- Extraneous Variables | Examples, Types & Controls
- Guide to Experimental Design | Overview, Steps, & Examples
- How Do You Incorporate an Interview into a Dissertation? | Tips
- How to Do Thematic Analysis | Step-by-Step Guide & Examples
- How to Write a Literature Review | Guide, Examples, & Templates
- How to Write a Strong Hypothesis | Steps & Examples
- Inclusion and Exclusion Criteria | Examples & Definition
- Independent vs. Dependent Variables | Definition & Examples
- Inductive Reasoning | Types, Examples, Explanation
- Inductive vs. Deductive Research Approach | Steps & Examples
- Internal Validity in Research | Definition, Threats, & Examples
- Internal vs. External Validity | Understanding Differences & Threats
- Longitudinal Study | Definition, Approaches & Examples
- Mediator vs. Moderator Variables | Differences & Examples
- Mixed Methods Research | Definition, Guide & Examples
- Multistage Sampling | Introductory Guide & Examples
- Naturalistic Observation | Definition, Guide & Examples
- Operationalization | A Guide with Examples, Pros & Cons
- Population vs. Sample | Definitions, Differences & Examples
- Primary Research | Definition, Types, & Examples
- Qualitative vs. Quantitative Research | Differences, Examples & Methods
- Quasi-Experimental Design | Definition, Types & Examples
- Questionnaire Design | Methods, Question Types & Examples
- Random Assignment in Experiments | Introduction & Examples
- Random vs. Systematic Error | Definition & Examples
- Reliability vs. Validity in Research | Difference, Types and Examples
- Reproducibility vs Replicability | Difference & Examples
- Reproducibility vs. Replicability | Difference & Examples
- Sampling Methods | Types, Techniques & Examples
- Semi-Structured Interview | Definition, Guide & Examples
- Simple Random Sampling | Definition, Steps & Examples
- Single, Double, & Triple Blind Study | Definition & Examples
- Stratified Sampling | Definition, Guide & Examples
- Structured Interview | Definition, Guide & Examples
- Survey Research | Definition, Examples & Methods
- Systematic Review | Definition, Example, & Guide
- Systematic Sampling | A Step-by-Step Guide with Examples
- Textual Analysis | Guide, 3 Approaches & Examples
- The 4 Types of Reliability in Research | Definitions & Examples
- The 4 Types of Validity in Research | Definitions & Examples
- Transcribing an Interview | 5 Steps & Transcription Software
- Triangulation in Research | Guide, Types, Examples
- Types of Interviews in Research | Guide & Examples
- Types of Research Designs Compared | Guide & Examples
- Types of Variables in Research & Statistics | Examples
- Unstructured Interview | Definition, Guide & Examples
- What Is a Case Study? | Definition, Examples & Methods
- What Is a Case-Control Study? | Definition & Examples
- What Is a Cohort Study? | Definition & Examples
- What Is a Conceptual Framework? | Tips & Examples
- What Is a Controlled Experiment? | Definitions & Examples
- What Is a Double-Barreled Question?
- What Is a Focus Group? | Step-by-Step Guide & Examples
- What Is a Likert Scale? | Guide & Examples
- What Is a Prospective Cohort Study? | Definition & Examples
- What Is a Retrospective Cohort Study? | Definition & Examples
- What Is Action Research? | Definition & Examples
- What Is an Observational Study? | Guide & Examples
- What Is Concurrent Validity? | Definition & Examples
- What Is Content Validity? | Definition & Examples
- What Is Convenience Sampling? | Definition & Examples
- What Is Convergent Validity? | Definition & Examples
- What Is Criterion Validity? | Definition & Examples
- What Is Data Cleansing? | Definition, Guide & Examples
- What Is Deductive Reasoning? | Explanation & Examples
- What Is Discriminant Validity? | Definition & Example
- What Is Ecological Validity? | Definition & Examples
- What Is Ethnography? | Definition, Guide & Examples
- What Is Face Validity? | Guide, Definition & Examples
- What Is Non-Probability Sampling? | Types & Examples
- What Is Participant Observation? | Definition & Examples
- What Is Peer Review? | Types & Examples
- What Is Predictive Validity? | Examples & Definition
- What Is Probability Sampling? | Types & Examples
- What Is Purposive Sampling? | Definition & Examples
- What Is Qualitative Observation? | Definition & Examples
- What Is Qualitative Research? | Methods & Examples
- What Is Quantitative Observation? | Definition & Examples
- What Is Quantitative Research? | Definition, Uses & Methods
What is your plagiarism score?
Red 15 Hz flickering light: a novel technique for effective wild bird management
- Published: 09 September 2024
- Volume 70 , article number 92 , ( 2024 )
Cite this article

- Takeshi Honda ORCID: orcid.org/0000-0002-3465-1907 1 ,
- Hiroki Tominaga 2 &
- Akio Shimizu 2
1 Altmetric
Human-bird conflicts are in a critical state, involving economic losses such as agricultural losses, bird strikes on aircraft and avian influenza. Traditional technologies leveraging bird vision and hearing often lose their effectiveness over time as birds become habituated to these stimuli. To address these challenges, our study introduces a novel countermeasure technology based on neurophysiology. The human brain reacts to flickering light, which can cause symptoms like headaches, nausea, and dizziness. In extremely rare cases, 15 Hz flickering red light can even lead to epilepsy. Not only humans, but chickens also suffer from 14 Hz flickering light. This led us to consider the possibility that similar flickering light stimuli could be applicable to bird management. In our experiments conducted during the day, we used long-range flashlights. White flickering light had no effect on bird escape behavior. However, when cellophane film was attached to the flashlights to restrict the wavelength, the emitted red light induced escape behavior in birds. Additionally, employing two types of flashlights to generate flickering red + blue or red + green lights elicited escape behavior. However, the blue and green combination proved to be less effective. The most intense flickering frequency for crows was 15 Hz. These results are highly similar to those found in human neurophysiology, showing that red light alone and the combination of red and blue lights have the most significant impact on the brain. By measuring the flight initiation distance (FID) of birds, we found that illuminated areas had a significantly higher FID (137 m) compared to non-illuminated areas (12 m). These findings suggest that applying principles of human physiology to wildlife management can offer new solutions for bird damage control.
This is a preview of subscription content, log in via an institution to check access.
Access this article
Subscribe and save.
- Get 10 units per month
- Download Article/Chapter or eBook
- 1 Unit = 1 Article or 1 Chapter
- Cancel anytime
Price includes VAT (Russian Federation)
Instant access to the full article PDF.
Rent this article via DeepDyve
Institutional subscriptions

Data availability
The data will be provided upon reasonable request to the corresponding author.
Avery ML, Genchi AC (2004) Avian perching deterrents on ultrasonic sensors at airport Wind-Shear Alert Systems. Wildl Soc Bull 32:718–725. https://doi.org/10.2193/0091-7648(2004)032[0718:APDOUS]2.0.CO;2
Article Google Scholar
Barnes TA, Dwyer JF, Mojica EK et al (2022) Wildland fires ignited by avian electrocutions. Wildl Soc Bull 46:e1302
Batini C, Teillet MA, Naquet R (2004) An avian model of genetic reflex epilepsy. Arch Ital Biol 142:297–312
PubMed CAS Google Scholar
Batra S, Pandav CS, Ahuja S (2019) Light emitting diode lighting flicker, its impact on health, and the need to minimise it. J Clin Diagnostic Res 13:1–5
Google Scholar
Blackwell BF, Bernhardt GE, Cepek JD, Dolbeer RA (2002) Lasers as non-lethal avian repellents: potential applications in the airport environment. Fed Aviat Adm Technol Transf Conf 2002:1–10
Blackwell BF, Bernhardt GE, Giuliano, (2004) Efficacy of aircraft landing lights in stimulating avoidance behavior in birds. J Wildl Manage 68:725–732. https://doi.org/10.2193/0022-541X(2004)068[0725:EOALLI]2.0.CO;2
Blackwell BF, DeVault TL, Seamans TW et al (2012) Exploiting avian vision with aircraft lighting to reduce bird strikes. J Appl Ecol 49:758–766
Bonnot NC, Hewison AJM, Morellet N et al (2017) Stick or twist: roe deer adjust their flight behaviour to the perceived trade-off between risk and reward. Anim Behav 124:35–46. https://doi.org/10.1016/j.anbehav.2016.11.031
Boström JE, Dimitrova M, Canton C et al (2016) Ultra-rapid vision in birds. PLoS ONE 11:e0151099
Article PubMed PubMed Central Google Scholar
Brown RN, Brown DH (2021) Robotic laser scarecrows: a tool for controlling bird damage in sweet corn. Crop Prot 146:105652
Clucas B, Marzluff JM, Mackovjak D, Palmquist I (2013) Do American crows pay attention to human gaze and facial expressions? Ethology 119:296–302
Conover MR (1984) Comparative effectiveness of avitrol, exploders, and hawk-kites in reducing blackbird damage to corn. J Wildl Manage 48:109–116
Conover MR (2001) Resolving Human-Wildlife Conflicts: The Science of Wildlife Damage Management. CRC Press, N.Y.
Book Google Scholar
Corcoran ME, Cain DP, Wada JA (1979) Photically induced seizures in the yellow baboon, Papio cynocephalus . Can J Neurol Sci 6:129–131
Article PubMed CAS Google Scholar
Da Silva AM, Leal B (2017) Photosensitivity and epilepsy: current concepts and perspectives—a narrative review. Seizure 50:209–218
Elbers ARW, Gonzales JL (2021) Efficacy of an automated laser for reducing wild bird visits to the free range area of a poultry farm. Sci Rep 11:12779
Article PubMed PubMed Central CAS Google Scholar
Fischer JW, Phillips GE, Baasch DM et al (2011) Modifying elk ( Cervus elaphus ) behavior with electric fencing at established fence-lines to reduce disease transmission potential. Wildl Soc Bull 35:9–14
Gentile CP, Aguirre GK (2020) A neural correlate of visual discomfort from flicker. J vis 20:11
Gilsdorf JM, Hygnstrom SE, VerCauteren KC et al (2004) Propane exploders and electronic guards were ineffective at reducing deer damage in cornfields. Wildl Soc Bull 32:524–531
Glahn JF, Ellis G, Fioranelli P, Dorr BS (2000) Evaluation of moderate and low-powered lasers for dispersing double-crested cormorants from their night roosts. Proc Ninth Wildl Damage Manage Conf 9:30–45
Gorenzel WP, Salmon TP (1995) Characteristics of American crow urban roosts in California. J Wildl Manage 59:638–645. https://doi.org/10.2307/3801939
Gorenzel WP, Blackwell BF, Simmons GD et al (2002) Evaluation of lasers to disperse American crows, Corvus brachyrhynchos , from urban night roosts. Int J Pest Manag 48:327–331. https://doi.org/10.1080/09670870210151689
Greenwood VJ, Smith EL, Goldsmith AR et al (2004) Does the flicker frequency of fluorescent lighting affect the welfare of captive European starlings? Appl Anim Behav Sci 86:145–159
Harding G (1998) TV can be bad for your health. Nat Med 4:265–267
Harding RM, Mills FJ (1983) Special forms of flight. II: helicopters. Br Med J 287:346–349
Article CAS Google Scholar
Hart NS (2001) The visual ecology of avian photoreceptors. Prog Retin Eye Res 20:675–703
Heim W, Piironen A, Heim RJ et al (2022) Effects of multiple targeted repelling measures on the behaviour of individually tracked birds in an area of increasing human-wildlife conflict. J Appl Ecol 59:3027–3037. https://doi.org/10.1111/1365-2664.14297
Hodos W (2003) Minimization of Motion Smear: Reducing Avian Collision with Wind Turbines
Honda T (2012) Line color affects the collision risk and deterrence of crows. J Ethol. https://doi.org/10.1007/s10164-011-0283-z
Honda T (2022) Height and tension of electric lines: how should an electric fence be installed to effectively mitigate human-wildlife conflict? Eur J Wildl Res 68:60. https://doi.org/10.1007/s10344-022-01606-6
Honda T, Ueda H (2023) Why mammals do not damage entire farmlands like insect pests do? A review from a behavioral perspective. Mammal Study 48:75–89
Kessler KK, Johnson RJ, Eskridge KM (1994) Monofilament lines and a hoop device for bird management at backyard feeders. Wildl Soc Bull 22:461–470
Knudsen EI (2020) Evolution of neural processing for visual perception in vertebrates. J Comp Neurol 528:2888–2901
Leblond M, Dussault C, Ouellet JP et al (2007) Electric fencing as a measure to reduce moose–vehicle collisions. J Wildl Manage 71:1695–1703
Lima SL, Valone TJ, Caraco T (1985) Foraging-efficiency-predation-risk trade-off in the grey squirrel. Anim Behav 33:155–165
Lovibond PF, Mitchell CJ, Minard E et al (2009) Safety behaviours preserve threat beliefs: Protection from extinction of human fear conditioning by an avoidance response. Behav Res Ther 47:716–720
Article PubMed Google Scholar
Maddocks SA, Goldsmith AR, Cuthill IC (2001) The influence of flicker rate on plasma corticosterone levels of European starlings, Sturnus vulgaris . Gen Comp Endocrinol 124:315–320
Manz ST, Sieving KE, Brown RN et al (2024) Experimental assessment of laser scarecrows for reducing avian damage to sweet corn. Pest Manag Sci 80:1547–1556
Mesri JC, Dellepiane C (1991) Colour and Photosensitive Epilepsy. Medicina (b Aires) 51:327–330
Mowrer OH, Lamoreaux RR (1946) Fear as an intervening variable in avoidance conditioning. J Comp Psychol 39:29–50
Parra J, da Silva FH, Stroink H, Kalitzin S (2007) Is colour modulation an independent factor in human visual photosensitivity? Brain 130:1679–1689
Peichl L (2005) Diversity of mammalian photoreceptor properties: adaptations to habitat and lifestyle? Anat Rec Adv Integr Anat Evol Biol 287:1001–1012
Potier S, Lieuvin M, Pfaff M, Kelber A (2020) How fast can raptors see? J Exp Biol 223:jeb209031
PubMed Google Scholar
Rash CE (2004) Awareness of causes and symptoms of flicker vertigo can limit ill effects. Hum Factors Aviat Med 51:1–6
Rebke M, Dierschke V, Weiner CN et al (2019) Attraction of nocturnally migrating birds to artificial light: The influence of colour, intensity and blinking mode under different cloud cover conditions. Biol Conserv 233:220–227
Reidinger JRF, Miller JE (2013) Wildlife Damage Management: Prevention, Problem Solving, and Conflict Resolution. JHU Press, Maryland
Reidy MM, Campbell TA, Hewitt DG (2008) Evaluation of electric fencing to inhibit feral pig movements. J Wildl Manage 72:1012–1018
Schiller PH (1992) The ON and OFF channels of the visual system. Trends Neurosci 15:86–92
Seamans TW, Lovell CD, Dolbeer RA, Cepek JD (2001) Evaluation of mirrors to deter nesting starlings. Wildl Soc Bull 29:1061–1066
Sharpe LT, Stockman A (1999) Rod pathways: the importance of seeing nothing. Trends Neurosci 22:497–504
Shirakawa S, Funatsuka M, Osawa M et al (2001) A study of the effect of color photostimulation from a Cathode-Ray Tube (CRT) display on photosensitive patients: The effect of alternating red-cyan flicker stimulation. Epilepsia 42:922–929
Souris M, Selenic D, Khaklang S et al (2014) Poultry farm vulnerability and risk of avian influenza re-emergence in Thailand. Int J Environ Res Public Health 11:934–951
Szabó CÁ, Fischer A (2021) Animal models of photosensitivity: clinical significance and windows into mechanisms. In: Kasteleijn-Nolst Trenite D (ed) The Importance of Photosensitivity for Epilepsy. Springer, Cham, pp 219–235
Chapter Google Scholar
Takahashi T, Tsukahara Y (1992) Usefulness of blue sunglasses in photosensitive epilepsy. Epilepsia 33:517–521
Takahashi T, Tsukahara Y, Kaneda S (1981) Influence of pattern and red color on the photoconvulsive response and the photic driving. Tohoku J Exp Med 133:129–137
Tedore C, Nilsson D-E (2019) Avian UV vision enhances leaf surface contrasts in forest environments. Nat Commun 10:238
Tierson WC (1969) Controlling deer use of forest vegetation with electric fences. J Wildl Manage 33:922–926
Tobimatsu S, Zhang YM, Tomoda Y et al (1999) Chromatic sensitive epilepsy: a variant of photosensitive epilepsy. Ann Neurol 45:790–793
Verdolin JL (2006) Meta-analysis of foraging and predation risk trade-offs in terrestrial systems. Behav Ecol Sociobiol 60:457–464
Walter VJ, Walter WG (1949) The central effects of rhythmic sensory stimulation. Electroencephalogr Clin Neurophysiol 1:57–86
Wang Z, Griffin AS, Lucas A, Wong KC (2019) Psychological warfare in vineyard: Using drones and bird psychology to control bird damage to wine grapes. Crop Prot 120:163–170
Wilkins AJ (2016) A physiological basis for visual discomfort: Application in lighting design. Light Res Technol 48:44–54
Wilkins A, Veitch J, Lehman B (2010) LED lighting flicker and potential health concerns: IEEE standard PAR1789 update. In: 2010 IEEE Energy Conversion Congress and Exposition. pp 171–178
Yu F, Wang X, Zhao Y, Li Z (2023) Influence of age, breeding state and approach direction on sensitivity to human gaze: a field study on Azure-winged magpies. Anim Cogn 26:1369–1379. https://doi.org/10.1007/s10071-023-01786-x
Download references
Acknowledgements
The authors would like to thank Kaori Muramatsu for her valuable assistance in organizing the data and supporting the research. During the preparation of this work, the author utilized GPT-4.0 to improve spelling and grammar.
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and affiliations.
Yamanashi Prefecture Agricultural Research Center, Shimoimai, Kai, 1100, Japan
Takeshi Honda
Yamanashi Industrial Technology Center, Otsu, Kofu, 2094, Japan
Hiroki Tominaga & Akio Shimizu
You can also search for this author in PubMed Google Scholar
Contributions
Conceptualization: TH; Flashlight development: HT, AS; Formal analysis and investigation: TH; Writing—original draft preparation: TH; Writing—review and editing: HT, AS; Resources: TH, HT, AS; Supervision: TH.
Corresponding author
Correspondence to Takeshi Honda .
Ethics declarations
Ethical approval.
All applicable international, national, and institutional guidelines for the care and use of animals were followed. This animal study was approved by the Yamanashi Prefectural Research Assessment Committee (#050901).
Competing interest
The authors declare no competing interests.
Additional information
Publisher's note.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (PDF 203 KB)
Supplementary file2 (jpg 1820 kb), rights and permissions.
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Reprints and permissions
About this article
Honda, T., Tominaga, H. & Shimizu, A. Red 15 Hz flickering light: a novel technique for effective wild bird management. Eur J Wildl Res 70 , 92 (2024). https://doi.org/10.1007/s10344-024-01846-8
Download citation
Received : 05 April 2024
Revised : 25 August 2024
Accepted : 30 August 2024
Published : 09 September 2024
DOI : https://doi.org/10.1007/s10344-024-01846-8
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Agriculture
- Bird strike
- Find a journal
- Publish with us
- Track your research
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- Dtsch Arztebl Int
- v.106(15); 2009 Apr
Types of Study in Medical Research
Bernd röhrig.
1 MDK Rheinland-Pfalz, Referat Rehabilitation/Biometrie, Alzey
Jean-Baptist du Prel
2 Zentrum für Präventive Pädiatrie, Zentrum für Kinder- und Jugendmedizin, Mainz
Daniel Wachtlin
3 Interdisziplinäres Zentrum Klinische Studien (IZKS), Fachbereich Medizin der Universität Mainz
Maria Blettner
4 Institut für Medizinische Biometrie, Epidemiologie und Informatik (IMBEI), Johannes Gutenberg Universität Mainz
The choice of study type is an important aspect of the design of medical studies. The study design and consequent study type are major determinants of a study’s scientific quality and clinical value.
This article describes the structured classification of studies into two types, primary and secondary, as well as a further subclassification of studies of primary type. This is done on the basis of a selective literature search concerning study types in medical research, in addition to the authors’ own experience.
Three main areas of medical research can be distinguished by study type: basic (experimental), clinical, and epidemiological research. Furthermore, clinical and epidemiological studies can be further subclassified as either interventional or noninterventional.
Conclusions
The study type that can best answer the particular research question at hand must be determined not only on a purely scientific basis, but also in view of the available financial resources, staffing, and practical feasibility (organization, medical prerequisites, number of patients, etc.).
The quality, reliability and possibility of publishing a study are decisively influenced by the selection of a proper study design. The study type is a component of the study design (see the article "Study Design in Medical Research") and must be specified before the study starts. The study type is determined by the question to be answered and decides how useful a scientific study is and how well it can be interpreted. If the wrong study type has been selected, this cannot be rectified once the study has started.
After an earlier publication dealing with aspects of study design, the present article deals with study types in primary and secondary research. The article focuses on study types in primary research. A special article will be devoted to study types in secondary research, such as meta-analyses and reviews. This article covers the classification of individual study types. The conception, implementation, advantages, disadvantages and possibilities of using the different study types are illustrated by examples. The article is based on a selective literature research on study types in medical research, as well as the authors’ own experience.
Classification of study types
In principle, medical research is classified into primary and secondary research. While secondary research summarizes available studies in the form of reviews and meta-analyses, the actual studies are performed in primary research. Three main areas are distinguished: basic medical research, clinical research, and epidemiological research. In individual cases, it may be difficult to classify individual studies to one of these three main categories or to the subcategories. In the interests of clarity and to avoid excessive length, the authors will dispense with discussing special areas of research, such as health services research, quality assurance, or clinical epidemiology. Figure 1 gives an overview of the different study types in medical research.

Classification of different study types
*1 , sometimes known as experimental research; *2 , analogous term: interventional; *3 , analogous term: noninterventional or nonexperimental
This scheme is intended to classify the study types as clearly as possible. In the interests of clarity, we have excluded clinical epidemiology — a subject which borders on both clinical and epidemiological research ( 3 ). The study types in this area can be found under clinical research and epidemiology.
Basic research
Basic medical research (otherwise known as experimental research) includes animal experiments, cell studies, biochemical, genetic and physiological investigations, and studies on the properties of drugs and materials. In almost all experiments, at least one independent variable is varied and the effects on the dependent variable are investigated. The procedure and the experimental design can be precisely specified and implemented ( 1 ). For example, the population, number of groups, case numbers, treatments and dosages can be exactly specified. It is also important that confounding factors should be specifically controlled or reduced. In experiments, specific hypotheses are investigated and causal statements are made. High internal validity (= unambiguity) is achieved by setting up standardized experimental conditions, with low variability in the units of observation (for example, cells, animals or materials). External validity is a more difficult issue. Laboratory conditions cannot always be directly transferred to normal clinical practice and processes in isolated cells or in animals are not equivalent to those in man (= generalizability) ( 2 ).
Basic research also includes the development and improvement of analytical procedures—such as analytical determination of enzymes, markers or genes—, imaging procedures—such as computed tomography or magnetic resonance imaging—, and gene sequencing—such as the link between eye color and specific gene sequences. The development of biometric procedures—such as statistical test procedures, modeling and statistical evaluation strategies—also belongs here.
Clinical studies
Clinical studies include both interventional (or experimental) studies and noninterventional (or observational) studies. A clinical drug study is an interventional clinical study, defined according to §4 Paragraph 23 of the Medicines Act [Arzneimittelgesetz; AMG] as "any study performed on man with the purpose of studying or demonstrating the clinical or pharmacological effects of drugs, to establish side effects, or to investigate absorption, distribution, metabolism or elimination, with the aim of providing clear evidence of the efficacy or safety of the drug."
Interventional studies also include studies on medical devices and studies in which surgical, physical or psychotherapeutic procedures are examined. In contrast to clinical studies, §4 Paragraph 23 of the AMG describes noninterventional studies as follows: "A noninterventional study is a study in the context of which knowledge from the treatment of persons with drugs in accordance with the instructions for use specified in their registration is analyzed using epidemiological methods. The diagnosis, treatment and monitoring are not performed according to a previously specified study protocol, but exclusively according to medical practice."
The aim of an interventional clinical study is to compare treatment procedures within a patient population, which should exhibit as few as possible internal differences, apart from the treatment ( 4 , e1 ). This is to be achieved by appropriate measures, particularly by random allocation of the patients to the groups, thus avoiding bias in the result. Possible therapies include a drug, an operation, the therapeutic use of a medical device such as a stent, or physiotherapy, acupuncture, psychosocial intervention, rehabilitation measures, training or diet. Vaccine studies also count as interventional studies in Germany and are performed as clinical studies according to the AMG.
Interventional clinical studies are subject to a variety of legal and ethical requirements, including the Medicines Act and the Law on Medical Devices. Studies with medical devices must be registered by the responsible authorities, who must also approve studies with drugs. Drug studies also require a favorable ruling from the responsible ethics committee. A study must be performed in accordance with the binding rules of Good Clinical Practice (GCP) ( 5 , e2 – e4 ). For clinical studies on persons capable of giving consent, it is absolutely essential that the patient should sign a declaration of consent (informed consent) ( e2 ). A control group is included in most clinical studies. This group receives another treatment regimen and/or placebo—a therapy without substantial efficacy. The selection of the control group must not only be ethically defensible, but also be suitable for answering the most important questions in the study ( e5 ).
Clinical studies should ideally include randomization, in which the patients are allocated by chance to the therapy arms. This procedure is performed with random numbers or computer algorithms ( 6 – 8 ). Randomization ensures that the patients will be allocated to the different groups in a balanced manner and that possible confounding factors—such as risk factors, comorbidities and genetic variabilities—will be distributed by chance between the groups (structural equivalence) ( 9 , 10 ). Randomization is intended to maximize homogeneity between the groups and prevent, for example, a specific therapy being reserved for patients with a particularly favorable prognosis (such as young patients in good physical condition) ( 11 ).
Blinding is another suitable method to avoid bias. A distinction is made between single and double blinding. With single blinding, the patient is unaware which treatment he is receiving, while, with double blinding, neither the patient nor the investigator knows which treatment is planned. Blinding the patient and investigator excludes possible subjective (even subconscious) influences on the evaluation of a specific therapy (e.g. drug administration versus placebo). Thus, double blinding ensures that the patient or therapy groups are both handled and observed in the same manner. The highest possible degree of blinding should always be selected. The study statistician should also remain blinded until the details of the evaluation have finally been specified.
A well designed clinical study must also include case number planning. This ensures that the assumed therapeutic effect can be recognized as such, with a previously specified statistical probability (statistical power) ( 4 , 6 , 12 ).
It is important for the performance of a clinical trial that it should be carefully planned and that the exact clinical details and methods should be specified in the study protocol ( 13 ). It is, however, also important that the implementation of the study according to the protocol, as well as data collection, must be monitored. For a first class study, data quality must be ensured by double data entry, programming plausibility tests, and evaluation by a biometrician. International recommendations for the reporting of randomized clinical studies can be found in the CONSORT statement (Consolidated Standards of Reporting Trials, www.consort-statement.org ) ( 14 ). Many journals make this an essential condition for publication.
For all the methodological reasons mentioned above and for ethical reasons, the randomized controlled and blinded clinical trial with case number planning is accepted as the gold standard for testing the efficacy and safety of therapies or drugs ( 4 , e1 , 15 ).
In contrast, noninterventional clinical studies (NIS) are patient-related observational studies, in which patients are given an individually specified therapy. The responsible physician specifies the therapy on the basis of the medical diagnosis and the patient’s wishes. NIS include noninterventional therapeutic studies, prognostic studies, observational drug studies, secondary data analyses, case series and single case analyses ( 13 , 16 ). Similarly to clinical studies, noninterventional therapy studies include comparison between therapies; however, the treatment is exclusively according to the physician’s discretion. The evaluation is often retrospective. Prognostic studies examine the influence of prognostic factors (such as tumor stage, functional state, or body mass index) on the further course of a disease. Diagnostic studies are another class of observational studies, in which either the quality of a diagnostic method is compared to an established method (ideally a gold standard), or an investigator is compared with one or several other investigators (inter-rater comparison) or with himself at different time points (intra-rater comparison) ( e1 ). If an event is very rare (such as a rare disease or an individual course of treatment), a single-case study, or a case series, are possibilities. A case series is a study on a larger patient group with a specific disease. For example, after the discovery of the AIDS virus, the Center for Disease Control (CDC) in the USA collected a case series of 1000 patients, in order to study frequent complications of this infection. The lack of a control group is a disadvantage of case series. For this reason, case series are primarily used for descriptive purposes ( 3 ).
Epidemiological studies
The main point of interest in epidemiological studies is to investigate the distribution and historical changes in the frequency of diseases and the causes for these. Analogously to clinical studies, a distinction is made between experimental and observational epidemiological studies ( 16 , 17 ).
Interventional studies are experimental in character and are further subdivided into field studies (sample from an area, such as a large region or a country) and group studies (sample from a specific group, such as a specific social or ethnic group). One example was the investigation of the iodine supplementation of cooking salt to prevent cretinism in a region with iodine deficiency. On the other hand, many interventions are unsuitable for randomized intervention studies, for ethical, social or political reasons, as the exposure may be harmful to the subjects ( 17 ).
Observational epidemiological studies can be further subdivided into cohort studies (follow-up studies), case control studies, cross-sectional studies (prevalence studies), and ecological studies (correlation studies or studies with aggregated data).
In contrast, studies with only descriptive evaluation are restricted to a simple depiction of the frequency (incidence and prevalence) and distribution of a disease within a population. The objective of the description may also be the regular recording of information (monitoring, surveillance). Registry data are also suited for the description of prevalence and incidence; for example, they are used for national health reports in Germany.
In the simplest case, cohort studies involve the observation of two healthy groups of subjects over time. One group is exposed to a specific substance (for example, workers in a chemical factory) and the other is not exposed. It is recorded prospectively (into the future) how often a specific disease (such as lung cancer) occurs in the two groups ( figure 2a ). The incidence for the occurrence of the disease can be determined for both groups. Moreover, the relative risk (quotient of the incidence rates) is a very important statistical parameter which can be calculated in cohort studies. For rare types of exposure, the general population can be used as controls ( e6 ). All evaluations naturally consider the age and gender distributions in the corresponding cohorts. The objective of cohort studies is to record detailed information on the exposure and on confounding factors, such as the duration of employment, the maximum and the cumulated exposure. One well known cohort study is the British Doctors Study, which prospectively examined the effect of smoking on mortality among British doctors over a period of decades ( e7 ). Cohort studies are well suited for detecting causal connections between exposure and the development of disease. On the other hand, cohort studies often demand a great deal of time, organization, and money. So-called historical cohort studies represent a special case. In this case, all data on exposure and effect (illness) are already available at the start of the study and are analyzed retrospectively. For example, studies of this sort are used to investigate occupational forms of cancer. They are usually cheaper ( 16 ).

Graphical depiction of a prospective cohort study (simplest case [2a]) and a retrospective case control study (2b)
In case control studies, cases are compared with controls. Cases are persons who fall ill from the disease in question. Controls are persons who are not ill, but are otherwise comparable to the cases. A retrospective analysis is performed to establish to what extent persons in the case and control groups were exposed ( figure 2b ). Possible exposure factors include smoking, nutrition and pollutant load. Care should be taken that the intensity and duration of the exposure is analyzed as carefully and in as detailed a manner as possible. If it is observed that ill people are more often exposed than healthy people, it may be concluded that there is a link between the illness and the risk factor. In case control studies, the most important statistical parameter is the odds ratio. Case control studies usually require less time and fewer resources than cohort studies ( 16 ). The disadvantage of case control studies is that the incidence rate (rate of new cases) cannot be calculated. There is also a great risk of bias from the selection of the study population ("selection bias") and from faulty recall ("recall bias") (see too the article "Avoiding Bias in Observational Studies"). Table 1 presents an overview of possible types of epidemiological study ( e8 ). Table 2 summarizes the advantages and disadvantages of observational studies ( 16 ).
| Study of rare diseases such as cancers | Case control studies |
| Study of rare exposure, such as exposure to industrial chemicals | Cohort studies in a population group in which there has been exposure (e.g. industrial workers) |
| Study of multiple exposures, such as the combined effect of oral contraceptives and smoking on myocardial infarction | Case control studies |
| Study of multiple end points, such as mortality from different causes | Cohort studies |
| Estimate of the incidence rate in exposed populations | Exclusively cohort studies |
| Study of covariables which change over time | Preferably cohort studies |
| Study of the effect of interventions | Intervention studies |
| Selection bias | N/A | 2 | 3 | 1 |
| Recall bias | N/A | 3 | 3 | 1 |
| Loss to follow-up | N/A | N/A | 1 | 3 |
| Confounding | 3 | 2 | 2 | 1 |
| Time required | 1 | 2 | 2 | 3 |
| Costs | 1 | 2 | 2 | 3 |
1 = slight; 2 = moderate; 3 = high; N/A, not applicable.
*Individual cases may deviate from this pattern.
Selecting the correct study type is an important aspect of study design (see "Study Design in Medical Research" in volume 11/2009). However, the scientific questions can only be correctly answered if the study is planned and performed at a qualitatively high level ( e9 ). It is very important to consider or even eliminate possible interfering factors (or confounders), as otherwise the result cannot be adequately interpreted. Confounders are characteristics which influence the target parameters. Although this influence is not of primary interest, it can interfere with the connection between the target parameter and the factors that are of interest. The influence of confounders can be minimized or eliminated by standardizing the procedure, stratification ( 18 ), or adjustment ( 19 ).
The decision as to which study type is suitable to answer a specific primary research question must be based not only on scientific considerations, but also on issues related to resources (personnel and finances), hospital capacity, and practicability. Many epidemiological studies can only be implemented if there is access to registry data. The demands for planning, implementation, and statistical evaluation for observational studies should be just as high for observational studies as for experimental studies. There are particularly strict requirements, with legally based regulations (such as the Medicines Act and Good Clinical Practice), for the planning, implementation, and evaluation of clinical studies. A study protocol must be prepared for both interventional and noninterventional studies ( 6 , 13 ). The study protocol must contain information on the conditions, question to be answered (objective), the methods of measurement, the implementation, organization, study population, data management, case number planning, the biometric evaluation, and the clinical relevance of the question to be answered ( 13 ).
Important and justified ethical considerations may restrict studies with optimal scientific and statistical features. A randomized intervention study under strictly controlled conditions of the effect of exposure to harmful factors (such as smoking, radiation, or a fatty diet) is not possible and not permissible for ethical reasons. Observational studies are a possible alternative to interventional studies, even though observational studies are less reliable and less easy to control ( 17 ).
A medical study should always be published in a peer reviewed journal. Depending on the study type, there are recommendations and checklists for presenting the results. For example, these may include a description of the population, the procedure for missing values and confounders, and information on statistical parameters. Recommendations and guidelines are available for clinical studies ( 14 , 20 , e10 , e11 ), for diagnostic studies ( 21 , 22 , e12 ), and for epidemiological studies ( 23 , e13 ). Since 2004, the WHO has demanded that studies should be registered in a public registry, such as www.controlled-trials.com or www.clinicaltrials.gov . This demand is supported by the International Committee of Medical Journal Editors (ICMJE) ( 24 ), which specifies that the registration of the study before inclusion of the first subject is an essential condition for the publication of the study results ( e14 ).
When specifying the study type and study design for medical studies, it is essential to collaborate with an experienced biometrician. The quality and reliability of the study can be decisively improved if all important details are planned together ( 12 , 25 ).
Acknowledgments
Translated from the original German by Rodney A. Yeates, M.A., Ph.D.
Conflict of interest statement
The authors declare that there is no conflict of interest in the sense of the International Committee of Medical Journal Editors.
- Open access
- Published: 04 September 2024
Genome-wide identification of the phenylalanine ammonia-lyase gene from Epimedium Pubescens Maxim. (Berberidaceae): novel insight into the evolution of the PAL gene family
- Chaoqun Xu 1 na1 ,
- Xuelan Fan 1 , 2 na1 ,
- Guoan Shen 1 &
- Baolin Guo 1
BMC Plant Biology volume 24 , Article number: 831 ( 2024 ) Cite this article
Metrics details
Phenylalanine ammonia-lyase (PAL) serves as a key gateway enzyme, bridging primary metabolism and the phenylpropanoid pathway, and thus playing an indispensable role in flavonoid, anthocyanin and lignin biosynthesis. PAL gene families have been extensively studied across species using public genomes. However, a comprehensive exploration of PAL genes in Epimedium species, especially those involved in prenylated flavonol glycoside, anthocyanin, or lignin biosynthesis, is still lacking. Moreover, an in-depth investigation into PAL gene family evolution is warranted.
Seven PAL genes ( EpPAL1 - EpPAL7 ) were identified. EpPAL2 and EpPAL3 exhibit low sequence identity to other EpPALs (ranging from 61.09 to 64.38%) and contain two unique introns, indicating distinct evolutionary origins. They evolve at a rate ~ 10 to ~ 54 times slower compared to EpPAL1 and EpPAL4-7 , suggesting strong purifying selection. EpPAL1 evolved independently and is another ancestral gene. EpPAL1 formed EpPAL4 through segmental duplication, which lead to EpPAL5 and EpPAL6 through tandem duplications, and EpPAL7 through transposed duplication, shaping modern EpPALs . Correlation analysis suggests EpPAL1 , EpPAL2 and EpPAL3 play important roles in prenylated flavonol glycosides biosynthesis, with EpPAL2 and EpPAL3 strongly correlated with both Epimedin C and total prenylated flavonol glycosides. EpPAL1 , EpPAL2 and EpPAL3 may play a role in anthocyanin biosynthesis in leaves. EpPAL2 , EpPAL3 , EpPAL6 , and EpPAL7 might be engaged in anthocyanin production in petals, and EpPAL2 and EpPAL3 might also contribute to anthocyanin synthesis in sepals. Further experiments are needed to confirm these hypotheses. Novel insights into the evolution of PAL gene family suggest that it might have evolved from a monophyletic group in bryophytes to large-scale sequence differentiation in gymnosperms, basal angiosperms, and Magnoliidae. Ancestral gene duplications and vertical inheritance from gymnosperms to angiosperms likely occurred during PAL evolution. Most early-diverging eudicotyledons and monocotyledons have distinct histories, while modern angiosperm PAL gene families share similar patterns and lack distant gene types.
Conclusions
EpPAL2 and EpPAL3 may play crucial roles in biosynthesis of prenylated flavonol glycosides and anthocyanins in leaves and flowers. This study provides novel insights into PAL gene family evolution. The findings on PAL genes in E. pubescens will aid in synthetic biology research on prenylated flavonol glycosides production.
Peer Review reports
Introduction
Phenylalanine ammonia-lyase (PAL, EC 4.3.1.24) is the first critical enzyme in the phenylpropanoid pathway, catalyzing the biotransformation of L-phenylalanine to trans-cinnamic acid. Acting as a bridge, PAL mediates carbon flux from primary to secondary metabolism, leading to the production of flavonoids, anthocyanins, tannins, lignins, phytoalexins and other benzene-based compounds with pharmaceutical value [ 1 , 2 ]. The phenylpropanoid derivatives play crucial roles in plant defense against a range of biotic (e.g., insects and pathogens) and abiotic stresses (e.g., UV light, low temperature and nutrient stress). These compounds function as regulatory molecules, participating in signal transduction and communication with other organisms [ 3 ]. Furthermore, they contribute to lignin biosynthesis, which is essential for maintaining stem rigidity, vascular integrity, and providing a physical barrier against invading pathogens in plants [ 4 , 5 ].
PAL enzymes in dicots typically exhibit monofunctionality, specifically catalyzing the PAL reaction. However, in certain monocots, particularly those belonging to the grass family Poaceae, PAL enzymes can display bifunctionality, catalyzing both PAL and TAL reactions with phenylalanine and tyrosine as substrates, respectively. Notably, PAL and TAL enzymes are absent in animals, where they have been replaced by HAL (L-histidine ammonia-lyase) [ 2 , 6 , 7 ]. The PAL gene family typically consists of 2–6 copies, although some species possess significantly more members [ 8 , 9 ]. Over the course of evolution, the expression of PAL genes in response to biotic and abiotic stresses has become highly regulated in a temporal and spatial manner, often resulting in the diversification of gene copies with redundant functions [ 10 , 11 , 12 ]. Given the diverse functions of PAL gene copies, it can be challenging to determine which copy predominantly modulates the biosynthesis of different branch end-products.
Herba Epimedii, a valued traditional Chinese medicine (TCM), is sourced exclusively from the dried leaves of four Epimedium species: namely E. pubescens , E. sagittatum , E. brevicornu and E. koreanum . Besides its traditional uses as a kidney tonic and antirheumatic agent [ 13 , 14 ], the aglycone of its primary bioactive components, known as prenylated flavonol glycosides (PFGs), particularly icaritin, has garnered recognition as a novel drug effective in inhibiting hepatocellular carcinoma (HCC) initiation and malignant growth [ 15 , 16 ]. However, the genes involved in PFGs biosynthesis in Epimedium , including PAL, remain fragmented. To date, only three PALs ( EsPAL1 , EsPAL2 and EsPAL3 ) in E. sagittatum [ 17 ] and one PAL ( EwPAL ) in E. wushanense [ 18 ] have been reported. Through qRT-PCR and correlation analysis, only EsPAL3 has been implicated in both PFGs and anthocyanin pathways. Meanwhile, EsPAL1 is suspected to play a role in lignin biosynthesis, while EsPAL2 demonstrates constitutive expression across all tissues, hinting at its potential involvement in lignin, PFGs and anthocyanin pathways. EwPAL , on the other hand, has been solely linked to the biosynthesis of naringenin. Further research into the PALs responsible for PFGs, anthocyanin or lignin biosynthesis needs to be clearly explored, which would facilitate more efficient synthesis of PFGs.
Previous studies on the evolution of the PAL gene family are limited, with only a few notable investigations reported: in Nelumbo nucifera by Wu et al. (2014) [ 19 ] and in three Cucurbitaceae plants by Dong et al. (2016) [ 8 ]. While the former study identified a distinct ancient NnPAL1 gene in the early-diverging dicotyledonous plant N. nucifera , it failed to explore similar patterns in other early-diverging dicotyledonous species. On the other hand, the latter study exclusively examined PAL evolution within Cucurbitaceae plants without delving into the evolutionary origins of the discovered PAL genes. E. pubescens , as another early-diverging dicotyledonous plant, emerges as a valuable subject for studying the evolution of the PAL gene family. Therefore, conducting in-depth research on this species becomes particularly significant.
In this study, a genome-wide search of E. pubescens led to the identification of 7 PAL genes. Among these, EpPAL2 and EpPAL3 exhibit significant sequence divergences, yet there is a lack of research exploring their distinct functional traits. This study aims to delve deeper into the gene functions of each EpPAL , with a particular focus on EpPAL2 and EpPAL3 . Furthermore, by utilizing E. pubescens as a representative of an early-diverging angiosperm, we aim to integrate prior research and employ a comprehensive set of 24 representative species to thoroughly examine the evolutionary history of the PAL gene family. The findings of this study have implications for deepening our understanding of the molecular functions of various EpPALs and providing valuable insights into the evolution of the PAL gene family.
Identification and chromosomal localization of EpPAL s
Putative PAL genes were retrieved and identified from E. pubescens by HMM search and BLASTP. A total of 7 PAL genes were identified. The full length of candidate genes was further confirmed to be correct using available transcriptome data of E. pubescens . According to the subcellular localization predictions, all EpPALs are in the cytoplasm. Detailed information of those genes is presented in Table 1 , including gene IDs, gene names, exon numbers, chromosome locations and protein length. The naming of EpPAL s was done according to the order of PAL on the chromosome of E. pubescens . 7 EpPALs were unevenly distributed across the whole genome. Specifically, EpPAL1 was located on chromosome 1, EpPAL2 and EpPAL3 were on chromosome 4, while EpPAL4 - EpPAL7 were allocated on chromosome 6 (Table 1 ).
Evolutionary analysis of PAL genes
To gain a deeper understanding of the evolution of PAL genes, we conducted a comprehensive analysis utilizing a diverse set of PAL members from 24 species, including Chara braunii (a charophyte), Physcomitrium patens (a bryophyte), Ceratopteris richardii (a fern), Ginkgo biloba and Sequoiadendron giganteum (gymnosperms), Amborella trichopoda and Nymphaea colorata (basal angiosperms), Ceratophyllum demersum (Ceratophyllales), Cinnamomum kanehirae , Liriodendron chinense and Piper nigrum (Magnoliidae), Spirodela polyrhiza (a basal monocot), Papaver somniferum , N. nucifera , Tetracentron sinense , Macadamia integrifolia , Aquilegia coerulea , Kingdonia uniflora and E. pubescens (early-diverging eudicotyledons), Brachypodium distachyon (an early-diverging monocotyledon), Oryza sativa (a representative modern monocotyledon), as well as A. thaliana , Cucumis sativus and Vitis vinifera (representative modern dicotyledons). Detailed gene mining instructions and sequences of the PAL genes from these 24 species are provided in Table S1 and Table S2 , respectively.
We constructed a phylogenetic tree using the maximum likelihood method to illustrate the differentiation profile of PAL genes among the aforementioned taxa (Fig. 1 ). As shown in Fig. 1 , the tree clearly divides all PAL genes into six distinct clusters (Cluster 1–6). Cluster 1 represents an evolutionary branch unique to charophytes, with a significantly longer branch length compared to other clusters, indicating that its PAL evolutionary clade is the farthest from the others. Cluster 2 and 4 exclusively contain gymnosperm genes, suggesting they may represent specific evolutionary branches unique to gymnosperms. Cluster 3 and 5 mainly include PAL genes from bryophytes, ferns, gymnosperms, basal angiosperms, Ceratophyllales, Magnoliidae, and early-diverging eudicotyledons, indicating a more primitive group of PALs. Cluster 6 primarily encompasses PAL clades from typical modern dicotyledons and monocotyledons, as well as homologous genes in ferns, gymnosperms, basal angiosperms, Magnoliidae, Ceratophyllales, and other taxa corresponding to modern PAL genes.
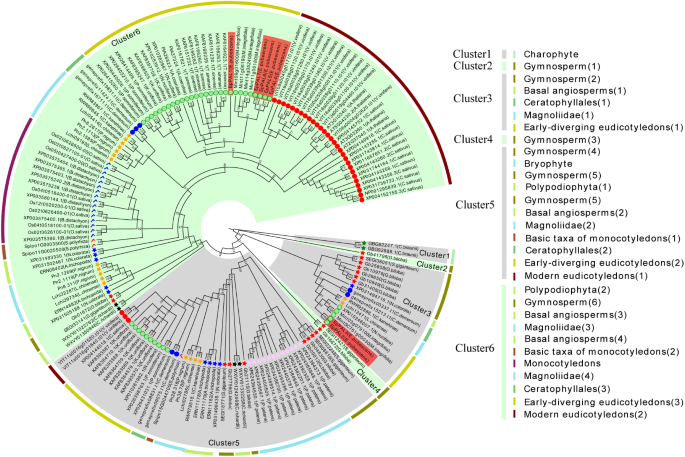
Phylogenetic tree of PAL sequences of 24 plant species with typical evolutionary relationships. EpPALs are marked with red box. The representation of the markings at the branch nodes is as follows: green five-pointed star: charophyte, red five-pointed star: gymnosperm, blue five-pointed star: basal angiosperms, orange five-pointed star: Magnoliidae, pink five-pointed star: bryophyte, black five-pointed star: polypodiophyta, green circle: early differentiated angiosperms, blue circle: Ceratophyllales, red circle: modern eudicotyledons, red checkmark: basic taxa of monocotyledons, and blue checkmark: monocotyledons. The attributes of each gene in the six clusters represented on the right side, and the numbers in parentheses represent branches of corresponding attributes. Taking gymnosperm as an example, it can be subdivided into six branches, represented by Gymnosperm (1) to Gymnosperm (6) respectively. Detailed species source information of PAL, as well as sequences can be referred to Table S1 and Table S2 , respectively
Based on these 12 different evolutionary branches (charophyte, bryophyte, fern, gymnosperms, basal angiosperms, Ceratophyllales, Magnoliidae, basic taxa of eudicotyledon, early-diverging eudicotyledons, an early-diverging monocotyledon, a representative modern monocotyledon, and representative modern dicotyledons), we can roughly outline the differentiation profile of PAL genes among various taxonomic groups. The PAL genes of charophytes and bryophytes each form a monophyletic group, respectively. Ferns exhibit two major sequence divergences. Significant differentiation occurred in gymnosperm, resulting in five branches: two unique to gymnosperms, two shared by some primitive taxa and early-diverging angiosperm ancestors, and one clustering with modern taxa. Both basal angiosperms and Magnoliidae also show substantial differentiation. Most early-diverging angiosperms, including Epimedium , and ancestral monocotyledon taxa possess ancestral primitive group genes. Modern monocotyledons and dicotyledons, represented by O. sativa and A. thaliana , mostly form monophyletic groups, and most of them do not have ancestral group genes. Thus, the evolution of PAL genes has progressed from a monophyletic group in bryophyte with small-scale functional differentiation to large-scale sequence differentiation in gymnosperms, basal angiosperms, and Magnoliidae. Most early-diverging eudicotyledons and monocotyledons have different evolutionary histories, while modern angiosperm taxa tend to be monophyletic with few ancestral group genes.
Among the seven EpPALs identified in our study, two genes ( EpPAL2 and EpPAL3 ) cluster together with the ancestral type PAL genes represented by Cluster 3, supported by high bootstrap values. The remaining five genes ( EpPAL1 and EpPAL4 - 7 ) form a separate monophyletic group (Cluster 6) with equally strong bootstrap support (Fig. 1 ). These findings suggest that the PAL genes in Epimedium originated from at least two ancestral PAL homologous genes. The presence of EpPALs in different branches of the phylogenetic tree indicates their diverse evolutionary histories and potential functional diversification within the species.
Gene structure analysis of PAL genes
Statistical analysis of 167 PAL genes from 24 species primarily revealed three distinct intron insertion patterns. Pattern 1 (59/167) lacked introns, Pattern 2 (65/167) had a single intron with a shorter front-end exon (~ 400 bp) compared to the back-end (~ 1750 bp), and Pattern 3 (7/167) contained two introns with exon lengths of ~ 1140 bp, ~ 540 bp, and ~ 540 bp, respectively (Figure S1 ). Notably, in E. pubescens , intron lengths varied widely, from 422 bp in EpPAL5 to 2862 bp in EpPAL4 , while exon lengths were highly conserved. EpPAL2 and EpPAL3 followed Pattern 3, but differed in intron phase: EpPAL2 had two phase 0 introns, whereas EpPAL3 had one phase 2 and one phase 0 intron. Both genes had conserved glutamine codon (CAG) at the second exon/intron boundary. All other EpPALs exhibited Pattern 2, with conserved intron insertion sites between the second and third bases of specific codons: arginine (CGA) for EpPAL1 , EpPAL6 , and EpPAL7 , and isoleucine (AUU) for EpPAL4 and EpPAL5 (Fig. 2 b and Figure S1 ). These differences suggest independent origins for the intron insertion events in these two gene sets.
Using a fungal PAL gene from Neurospora tetrasperma (NCBI accession number: EGZ69514.1) as an outgroup, we determined the root position of the phylogenetic tree composed of EpPALs to infer their lineage. The research results support the division of EpPALs into two major clades, with Clade1 being further subdivided into two sub-clades (clade1_1 and clade1_2). Detailed homology detection and structural alignment among EpPALs were provided (Fig. 2 b and Table S3 ). Pairwise identity between EpPALs (excluding EpPAL2 and EpPAL3 ) ranged from 85.10 to 100%, indicating close relatedness. However, protein identity between Clade2 and Clade1 was lower, ranging from 61.09 to 64.38% (Fig. 2 a and Table S3 ), suggesting multiple ancestral origins for the formation of EpPALs . Phylogenetic analysis (Fig. 1 ), sequence alignment (Table S3 ) and gene structure (Fig. 2 b) collectively provide compelling evidence supporting the hypothesis that EpPAL2 and EpPAL3 trace their origins to a unique ancestral gene, whereas the other EpPALs descended from a different ancestral gene.
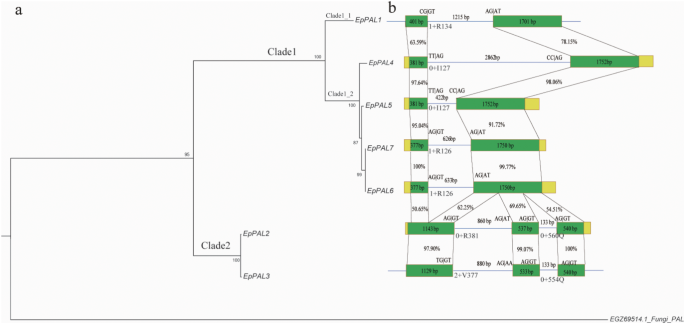
Phylogenetic tree and genomic structure of EpPALs . ( a ) Phylogenetic tree of EpPALs . The numbers below the branches represent the bootstrap values; ( b ) Gene structure of EpPALs . Green boxes, lines and yellow boxes represent the exon, intron and UTR, respectively. The percentages indicate the similarity of fragments between each pair of EpPALs . The exon/intron borders were displayed on top of the structural model, the intron phase (the numbers 0, 1 and 2, which represent introns between codons, introns between the first and second bases of a codon, introns between the second and the third bases of a codon, respectively), the amino acid residues affected by intron insertion events and its position were displayed under the structural model
Collinearity analysis of PALs
To determine the origin of EpPALs through duplication events, both inter- and intra-specific collinearity analyses were conducted (Fig. 3 ). We selected three species from different evolutionary branches ( A. trichopoda , N. nucifera , and O. sativa ) along with E. pubescens for microsynteny analysis of interspecies local regions related to PAL genes. Collinear blocks were detected for EpPAL1 to EpPAL6 , except for EpPAL7 . The analysis of EpPAL1 , EpPAL2 and EpPAL3 revealed collinear blocks only among E. pubescens , A. trichopoda , and N. nucifera , with no collinear block detected in O. sativa . Notably, homologous genes corresponding to EpPAL2 and EpPAL3 were not detected in A. trichopoda . Considering the detection of collinear blocks for EpPAL1 - 3 only in relatively primitive evolutionary branches and not in more modern species, it is speculated that the EpPAL1 - 3 may represent primitive types of the PAL family in E. pubescens . Further referencing Fig. 1 , we infer that the ancestral genes of EpPAL2 and EpPAL3 originated differently from that of EpPAL1 . The ancestral genes of EpPAL2 and EpPAL3 belong to Clade 3, a more primitive branch, while the ancestral gene of EpPAL1 belongs to Cluster 6, a branch present in modern monocots and dicots. For EpPAL4 - 6 , we detected relevant collinear blocks in species from three different evolutionary branches. Combining this with the evolutionary positions of these three genes in Fig. 1 , we infer that they may have originated from gene duplication of either the EpPAL1 branch or the EpPAL2 - 3 branch.
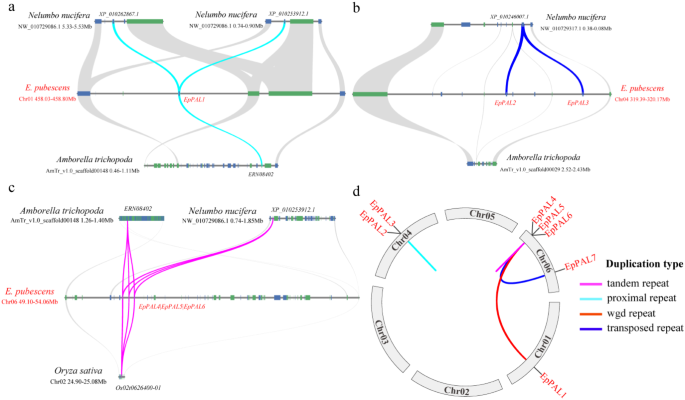
Inter- and intra-specific collinearity analysis of EpPALs . ( a ) Microsynteny analysis between the EpPAL1 loci and their respective collinear counterparts in A.trichopoda and N. nucifera . The collinear PAL genes and the syntenic flanking genes are connected by colored and gray lines, respectively; ( b ) Microsynteny analysis of EpPAL2 and EpPAL3 ; ( c ) Microsynteny analysis of EpPAL4 , EpPAL5 and EpPAL6 ; ( d ) Intraspecific collinearity analysis of EpPALs. Red line represents a collinear gene pair of EpPALs (the two ends were EpPAL1 and EpPAL4 , respectively). Purple line represents a tandem repeat between EpPAL4 and EpPAL5 - 6 . Cyan line represents a proximal repeat between EpPAL2 and EpPAL3 . Blue line represents a transposed repeat between EpPAL4 and EpPAL7
To gain a deeper understanding of the relationships among these genes, we conducted further intraspecific collinearity analysis in E. pubescens and analyzed gene duplication patterns using DupGen-finder (Fig. 3 d and Table S4 ). The results indicated that the gene pair EpPAL1 and EpPAL4 underwent segmental/whole-genome duplication (Fig. 3 d). EpPAL5 and EpPAL6 originated from tandem gene duplication of EpPAL4 , while EpPAL7 emerged from a transposed gene duplication event involving EpPAL4 (Fig. 3 d, Figure S2 and Table S4 ). Therefore, duplication events have played a significant role in the evolution of EpPALs . The evolutionary profile of EpPALs can be summarized as follows: the ancestral genes of E. pubescens are EpPAL2 and EpPAL3 , with EpPAL1 originating independently. EpPAL4 was then acquired through intraspecific whole-genome duplication from EpPAL1 . EpPAL4 underwent tandem duplication to produce EpPAL5 and EpPAL6 , and transposition duplication to generate EpPAL7 .
Conserved motif identification and cis- regulatory elements analysis
Eight conserved motifs followed a consistent distribution pattern of 6-4-7-3-8-1-2-5 among all EpPAL proteins (Figure S3 ). These motifs, ranging from 26 to 100 amino acids in length, were identified across all EpPALs (Figure S3 and Table S5 ). Notably, the MIO (4-methylidene-imidazolone-5-one) domain, characterized by the highly conserved signature sequence GTITASGDLVPLSYIA, contained the enzymatic active site Ala-Ser-Gly, which was preserved in all EpPAL proteins (Figure S3 ). With the exception of EpPAL3, which lacked the active site 158 L, the 388 F substrate-selective binding site, and the 350R phosphorylation site, both the active sites and substrate-specific binding sites were conserved across all EpPAL proteins (Figure S4 ). Additionally, while the 538 S phosphorylation site was serine in EpPAL2, EpPAL3 and EpPAL4, it was replaced by threonine in the remaining EpPAL proteins.
A total of 56 cis -regulatory elements (CREs) with known functions were identified, including 17, 19, 12, and 8 CREs related to light, stress, hormone, growth, and development responses, respectively (Fig. 4 ). With the exception of EpPAL4 , all EpPAL genes possess at least one AC element essential for lignin synthesis. However, no MBSI element related to the regulation of flavonoid biosynthetic genes was found. ARE (anaerobic induction), ABRE (abscisic acid-responsiveness), CGTCA-motif, and TGACG-motif (MeJA-responsiveness) were identified in all EpPAL genes, suggesting that responses to oxygen, abscisic acid, and methyl jasmonate are essential functions shared by all EpPALs . Additionally, ERE (ethylene-responsiveness) was exclusively present in EpPAL1 , while the CCGTCC-box was only found in EpPAL4 . These findings suggest that EpPAL1 and EpPAL4 may be specifically associated with ethylene response and activation of the meristem, respectively. Overall, these results imply that different EpPAL genes may have distinct yet overlapping biological functions, playing roles in processes such as growth and development, hormone response, and environmental stress response.
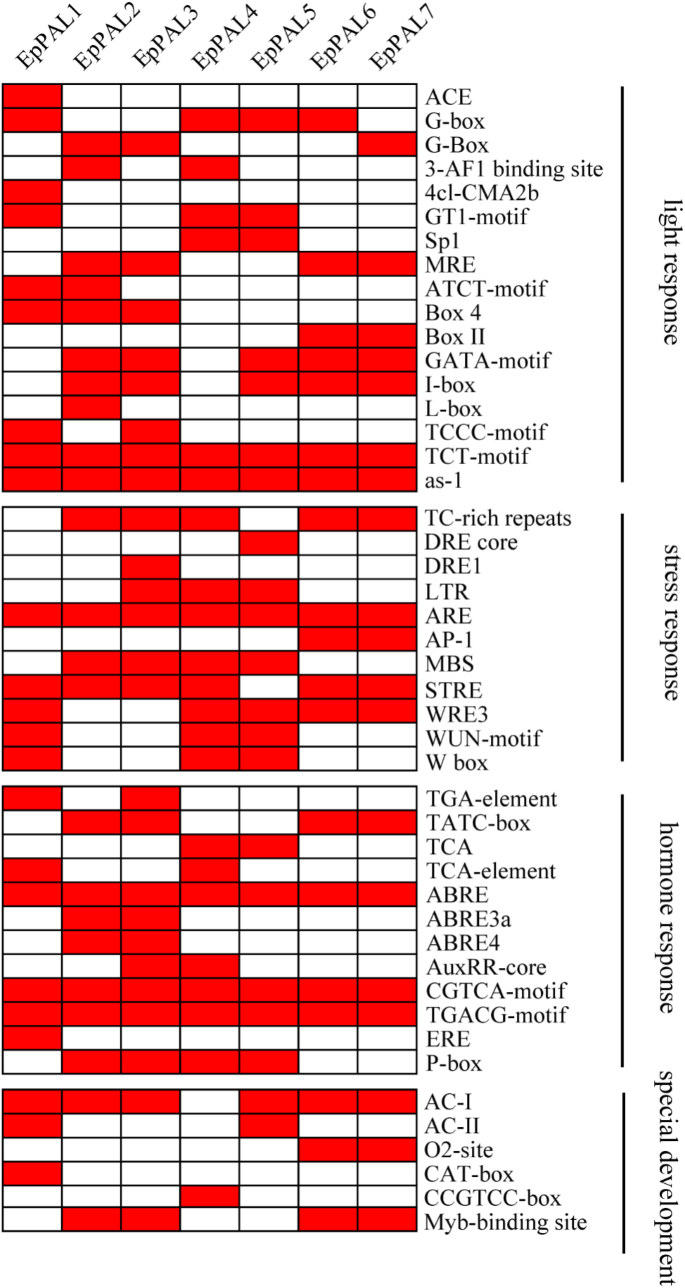
Cis -regulatory elements of EpPAL s in upstream region of 1500 bp. Four different types of cis -regulatory elements and different cis -regulatory elements of all EpPALs were provided
Natural selection analysis
Natural selection tests were conducted using PAML (v.4.1) under different hypotheses. For the branch-specific model, four hypotheses were tested as outlined in Table S6 . The three-ratio model, which assigns distinct ω values to each of the three clades (ω[Clade1_1] ≠ ω[Clade1_2] ≠ ω[Clade2]), emerged as a significantly better fit to the dataset compared to the other models tested (df = 1, P = 1.13e-07). Notably, the two-ratio model also outperformed the one-ratio model (df = 1, P = 2.62e-08), as illustrated in Fig. 2 a and detailed in Table S6 . These findings suggest that each clade experienced unique selection pressures, with ω ratios of 0.20, 0.037, and 0.0037 for Clade1_1, Clade1_2, and Clade2, respectively (Fig. 2 a and Table S6 ). Notably, Clade2 exhibited strong purifying selection, evolving approximately 54 and 10 times slower than Clade1_1 and Clade1_2, respectively. Clade1_2 followed in terms of purifying selection, while EpPAL1 , originating from Clade1_1, exhibited relatively higher divergence.
To further investigate whether ω varied across all amino acid sites or specific sites within particular branches, both the site-specific model and the branch-site model were employed. Under the site-specific model, three amino acid residues under positive selection when comparing selection M1 versus neutral M1, as well as M7 versus M8. Additionally, when setting Clade1_1, Clade1_2, and Clade2 as the foreground branches, 8, 7, and 76 amino acid sites were found under positive selection, respectively (Table S6 ). These candidate positively selected sites provide valuable insights into the evolutionary history of EpPALs .
Expression patterns of EpPALs and determination of EpPALs related to prenylated flavonol glycosides
The EpPALs were identified in five distinct tissues, as depicted in Figure S5 , with varying expression patterns across these tissues. EpPAL1 demonstrated high expression levels in all tested tissues, peaking in the leaf and flower, and gradually decreasing in the root, fruit, and stem, suggesting a fundamental role. In contrast, EpPAL2 and EpPAL3 showed elevated expression in leaf but were barely detectable in the stem, while EpPAL5 was predominantly expressed in leaf and root. EpPAL4 , EpPAL6 , and EpPAL7 had minimal or low levels of expression.
To further investigate the relationship between EpPAL expression and the contents of key bioactive compounds, a transcriptome analysis using samples from five different leaf development stages. The findings, along with the corresponding EpPAL expression levels and bioactive compound contents, are outlined in Table S7 . Additionally, a correlation analysis was performed, and the results are visually presented in Fig. 5 .
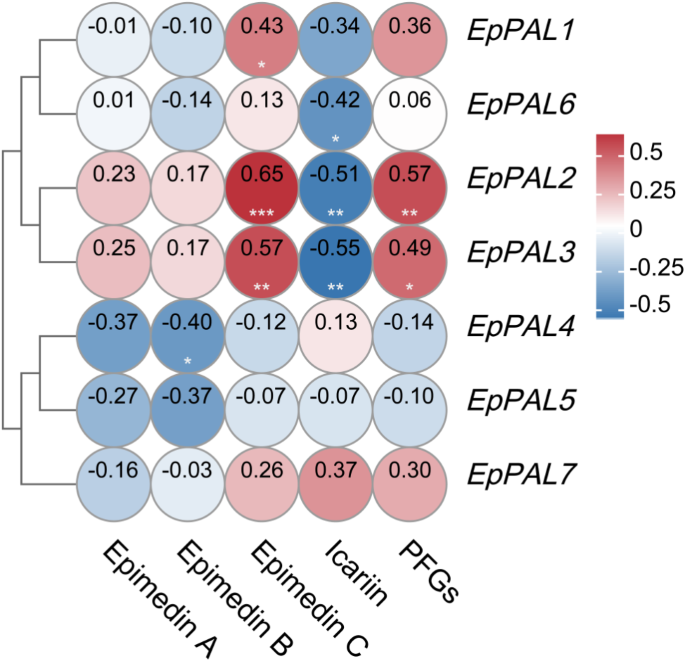
Correlation heatmap of EpPAL s with Epimedin A , Epimedin B , Epimedin C , Icariin and total PFGs. The significance levels were set as follows: unmarked stands for P > = 0.05, * stands for 0.01 < P < 0.05, ** stands for 0.001 < P < 0.01, *** stands for P < = 0.001
By integrating transcriptome data with targeted metabolite measurements, significant correlations were discovered between the expression of EpPAL2 and EpPAL3 with Epimedin C ( EpPAL2 : r = 0.65, P < 0.001; EpPAL3 : r = 0.57, P < 0.01) and total PFGs ( EpPAL2 : r = 0.57, P < 0.01; EpPAL3 : r = 0.49, P < 0.05). Notably, EpPAL1 holds the third position in terms of correlation strength. However, a notable negative correlation was observed between these genes and Icariin ( EpPAL2 : r = -0.51, P < 0.01; EpPAL3 : r = -0.55, P < 0.01). The remaining EpPALs demonstrated either weak ( EpPAL6 ) or no correlation ( EpPAL4 , EpPAL5 and EpPAL7 ), implying that EpPAL1 , EpPAL2 , and EpPAL3 , particularly the latter two, might have a pivotal role in the biosynthesis of PFGs.
Determination of EpPALs related to anthocyanin biosynthesis in flowers and leaves
To further identity the EpPAL isoforms responsible for anthocyanin biosynthesis, various groups of Epimedium species with distinct petal, sepal, and leaf colors were studied (Fig. 6 a). In leaves, EpPAL1 - 3 align with the observed color phenotypes, with EpPAL2 and EpPAL3 showing significantly higher expression in magenta leaves compared to green leaves, while EpPAL1 did not exhibit a notably high expression pattern (Fig. 6 b). Co-expression analysis with EpANSs and EpDFRs revealed a significant positive correlation between EpPAL1 - EpPAL3 and the expression of DFR and ANS genes, suggesting their involvement in anthocyanin biosynthesis in leaves, whereas EpPAL6 and EpPAL7 showed a significant negative correlation (Fig. 6 e and Table S9 ).
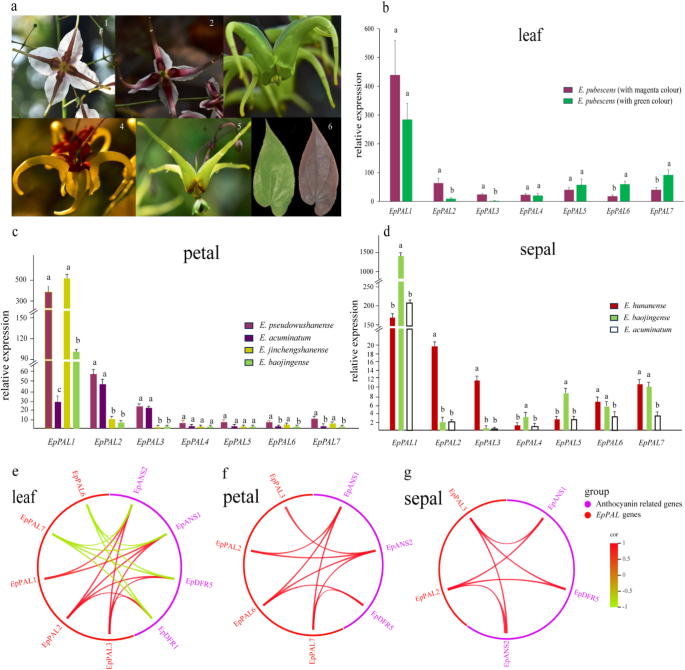
Phenotype of flower and leaf colors in different species of Epimedium plants, their PAL expression profiles, and the co-expression relationship with major anthocyanin-related genes. ( a ) Different colors in leaf, petal and sepal. a1 ~ a5 showed E. pseudowushanense , E. acuminatum , E. baojingense , E. hunanense and E. jinchengshanense , respectively. a6 showed leaf colors in E. pubescens with green and magenta, respectively. ( b ) Expression levels of EpPALs with different leaf colors. Significance tests for each EpPAL gene in two types of leaves were conducted and were labled with italicized ‘ a ’ and ‘ b ’ accordingly; ( c ) Expression levels of EpPALs in E. pseudowushanense , E. acuminatum , E. jinchengshanense and E. baojingense , with petal colors of magenta, magenta, yellow, and green, respectively. ( d ) Expression levels of EpPALs in E. hunanense , E. baojingense and E. acuminatum with sepal colors of red, green and white, respectively. Significance tests for each EpPAL gene in different colours of petals or sepals were conducted and were labeled with regular ‘a’ and ‘b’ accordingly. ( e-g ) Co-expression relationship between EpPALs and anthocyanin-related genes in leaf, petal and sepal, respectively. Three biological replicates were provided, and each repeat represent a mixing sample originated from three individuals
Similarly, in flowers, EpPAL2 and EpPAL3 showed significantly elevated expression in magenta petals compared to yellow petals and green petals (Fig. 6 c). Both genes exhibited similar expression patterns across different sepal colors (Fig. 6 d). Co-expression analysis in petals and sepals revealed a significant positive correlation between EpPAL2 , EpPAL3 , EpPAL6 and EpPAL7 with EpANSs and EpDFRs in petals (Fig. 6 f and g and Table S9 ), and between EpPAL2 and EpPAL3 with EpANSs and EpDFRs in sepals (Fig. 6 g and Table S9 ). Based on these expression profiles of EpPALs and co-expression patterns, we speculate that EpPAL2 and EpPAL3 may primarily be involved in anthocyanin synthesis in petals and sepals, while EpPAL6 and EpPAL7 may also play a role in anthocyanin synthesis in petals.
Repeatability verification of PFGs content dynamics and expression levels
To further validate the casual PAL genes involved in PFGs biosynthesis, a parallel validation experiment was conducted, comprising the determination of Epimedin C and total PFGs using the UPLC method (Fig. 7 c and Table S8 ), and the absolute quantification of seven EpPALs gene expressions across five developmental stages (S1 to S5) using the qRT-PCR method (Fig. 7 b). Six pairs of specific primers were designed for this purpose, with a single pair used for EpPAL6 and EpPAL7 due to their high sequence conservation (Table S10 ).
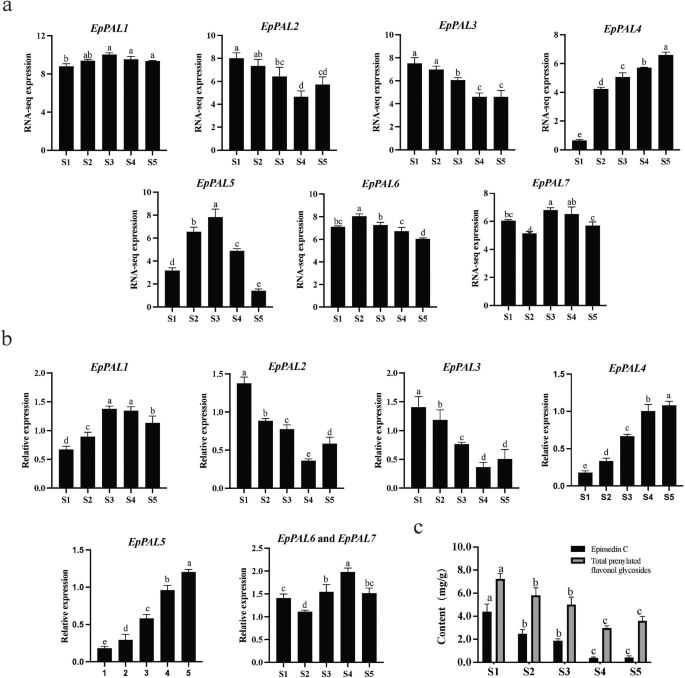
Expression patterns of EpPALs . ( a ) Expression patterns of EpPALs in S1-S5 by RNA-seq. The replicates can be referred to Table S7 . ( b ) Expression levels of EpPALs in S1-S5 by qRT-PCR. Three biological replicates were provided, and each repeat represent a mixing sample originated from three individuals. ( c ) The content of Epimedin C and total PFGs in S1-S5. S1-S5 represent the different stages of leaf development in E. pubescens . Error bars indicate standard error. The significance level is 0.05
The results revealed that EpPAL2 and EpPAL3 exhibited peak expression in S1, followed by a continuous decrease from S2 to S4, and a slight increase in S5 (Fig. 7 b), consistent with the relative quantification observed in transcriptome results (Fig. 7 a). A similar trend was observed for other EpPALs , except for EpPAL5 , indicating the overall reliability of the transcriptome data.
Importantly, the dynamic expression patterns of EpPAL2 and EpPAL3 across the five stages aligned with the dynamic changes in the content of Epimedin C and total PFGs (Fig. 7 ). However, no apparent correlation was found between the PFGs contents and qRT-PCR results for other EpPALs . For instance, EpPAL1 showed an initial increase (S1 to S3) followed by a decrease (S3 to S5), while EpPAL4 and EpPAL5 exhibited a continuous increasing trend. EpPAL6 and EpPAL7 fluctuated significantly, with the lowest expression in S2 and the highest in S4, respectively (Fig. 7 b), and did not align with the dynamic changes observed in Epimedin C and total PFGs content (Fig. 7 c).
Overall, our findings suggest that EpPAL2 and EpPAL3 may be the most critical genes responsible for the biosynthesis of PFGs in E. pubescens .
Characterization of the PAL gene family in E. pubescens
In the present study, the high-quality chromosome-level genome of E. pubescens served as a valuable resource for PAL research. A comprehensive exploration identified seven PAL genes ( EpPAL1 to EpPAL7 ), offering a more extensive analysis compared to previous studies in E. sagittatum [ 17 ]. All seven EpPALs were located in the cytoplasm, consistent with current PAL studies [ 8 , 9 , 12 , 19 ]. Despite the widely recognized conservation among EpPALs , two genes, EpPAL2 and EpPAL3 , exhibited lower identity (61.09–64.38%) to other EpPALs , reminiscent of ClPAL2 in Citrullus lanatus [ 20 ], which was proven to be an ancestral PAL, sharing only ~ 60% identity with other ClPALs despite having pairwise identities ranging from 71.2 to 99.0% [ 8 ]. Significant differences in PAL evolution with distinct origins were observed in E. pubescens , contributing to the varying sequence identities.
This study included a substantial proportion (20 out of 24) of primitive taxa which is different from previous studies [ 8 , 19 ]. This facilitated an investigation into intron insertion events in E. pubescens . While similar exon/intron structures tend to cluster together, as observed in E. sagittatum [ 17 ], pear [ 21 ], and common walnut [ 22 ], EpPAL2 and EpPAL3 gained two introns (Fig. 2 b and Figure S1 ), whereas other EpPALs possess only one intron insertion [ 8 , 9 , 11 , 12 , 17 , 23 , 24 ], suggesting significant structural divergence within the PAL gene family in E. pubescens .
Differences in intron/exon structures between EpPAL2-3 and other angiosperms were also detected (Figure S1 ), further indicating their ancient with distinct evolutionary origins. The clustering of the ancient gene NnPAL1 ( XP_010246007.1 in this study) with EpPAL2 and EpPAL3 supports this conclusion [ 19 ]. The increased intron number in ancient EpPAL genes may be significant, as intron-gain events can enhance mRNA stability or harbor regulatory elements without disrupting the coding frame of genes [ 25 , 26 , 27 , 28 ]. However, intron insertions in EpPAL2 , EpPAL3 and other EpPALs may have occurred independently based on their phylogenetic positions (Fig. 1 ).
Evolution of the plant PAL gene family
Previous studies have primarily focused on research at the species or family level [ 20 , 21 , 23 , 24 , 29 ], with limited investigations on the evolution of PAL genes, except for those in N. nucifera [ 19 ] and cucurbit species [ 8 ]. To bridge this gap, we conducted a phylogenomic study to elucidate the evolution of plant PAL (Fig. 1 ). Our findings reveal a large-scale gene duplication event in gymnosperms, with two gymnosperm-specific branches (cluster2 and cluster 4), emphasizing their unique origin. The presence of PALs in gymnosperms across other branches (cluster3, cluster5 and cluster6) suggests ancestral gene duplication and vertical inheritance during evolution, supporting widespread differentiation [ 19 ]. We hypothesize that the multigene families of PAL in gymnosperms may confer diverse functions, such as producing additional trans-cinnamic acid for downstream metabolic pathways or enhancing lignin biosynthesis to defend against adverse environments like insect and pathogen attacks [ 25 ], contributing to their widespread habitat adaptation.
Furthermore, our results indicate that the origin of PAL in angiosperm plants may not be monophyletic. This is evident from the evolutionary tree, where PAL genes of dicots like cucumber and lotus are clustered within cluster6 and also share clustering with PAL genes of primitive types within cluster3 and cluster5 (Fig. 1 ). This finding validates previous conclusions in N. nucifera [ 19 ]. Except for A. coerulea (only in cluster6), the PALs of other early-diverging eudicotyledons including E. pubescens (in cluster3 and 6), P. somniferum (in cluster5 and 6), M. integrifolia (in cluster3 and 6), K. uniflora (in cluster5 and 6), N. nucifera (in cluster3, 5 and 6), and T. sinense (cluster5 and 6) are divided into two clusters or three clusters ( N. nucifera ), with one clustering with cluster3 or cluster5, speculated to be relatively ancient genes, and the other with PALs of modern dicots (cluster6). This suggests different evolutionary origins may underlie the PAL gene evolution in early-diverging dicots [ 19 ]. Additionally, we speculate that ancient genes like EpPAL2 and EpPAL3 may have played a crucial role in the survival of these “pioneer species” in harsh environments during evolution, highlighting their importance for related plants. For instance, in E. pubescens , the positive selection test indicates that EpPAL2 and EpPAL3 experienced the strictest selection pressure, evolving ~ 10 and ~ 54 times slower than genes of Clade 1_2 ( EpPAL4 ~ 7 ) and Clade 1_1 ( EpPAL1 ), respectively (Table S6 ).
Expression profiles of EpPALs
Duplicate genes can lead to pseudogenization, subfunctionalization and neofunctionalization [ 2 ]. Gene expression patterns offer valuable insights into the functional differentiation. Previous research has shown that AtPAL1 and AtPAL2 in A. thaliana are highly expressed in roots and involved in stress-induced flavonoid biosynthesis [ 30 , 31 ]. In Pyrus bretschneideri , PbPAL1 and PbPAL2 are predominantly expressed in stems and roots, suggesting involvement in lignin synthesis and stone cell development [ 21 ].
In this study, the expression levels of EpPAL2 and EpPAL3 gradually decreased in correlation with the PFGs content during leaf development, indicating a potential significant role in PFGs biosynthesis. Our findings support the speculation that both genes positively correlate with Epimedin C and total PFGs content (Figs. 5 and 7 ). EpPAL1 may also be involved in PFGs biosynthesis in leaves, as evidenced by the third-highest correlation with Epimedin C and total PFGs (Fig. 5 ). As an ancestral gene with an independent origin, EpPAL1 is constitutively highly expressed across tissues, suggesting divergent functionality. Similar expression patterns of PAL can be observed in Cuminum cyminum and cucurbit plants [ 8 , 32 ]. Given the pivotal role of PAL in both primary and secondary metabolism, constitutive multifunctional expression may be crucial for maintaining diverse biological processes and adapting to changing environments.
In E. sagittatum , the expression patterns of EsPAL1 , EsPAL2 , and EsPAL3 align with lignification and active components accumulation [ 17 ]. In our study, EpPALs exhibited distinct yet overlapping expression patterns during different stages of leaf development (Fig. 5 and Table S7 ), hinting at possible functional diversification and redundancy. EpPAL2 and EpPAL3 demonstrated overlapping expression in PFGs and anthocyanin biosynthesis, ensuring functional redundancy, preventing the complete loss of weaker gene copies, a phenomenon often observed in key genes involved in different metabolic pathways [ 33 , 34 ]. In contrast, EpPAL1 and other EpPALs displayed distinct expression patterns, potentially stemming from functional differentiation among duplicate EpPAL genes, ultimately leading to their subfunctionalization and neofunctionalization.
In summary, this study comprehensively investigated PAL genes in E. pubescens . We speculate that EpPAL2 and EpPAL3 may participate in PFGs and anthocyanin pathways in leaves and flowers, while EpPAL1 , characterized by its constitutively high expression, may not only be involved in the biosynthesis of PFGs and anthocyanins in leaves, but also play a significant role in defense and protection. EpPAL6 and EpPAL7 may participate in anthocyanin synthesis in petals, but there is no evidence of their involvement in PFGs biosynthesis. Given the low expression levels, we hypothesize that EpPAL4 to EpPAL7 may function as stress responders or have become nonfunctional following duplication events. Further experimental validation is needed to confirm these speculations.
Materials and methods
Experimental materials.
The experimental materials were sourced from the Epimedium Germplasm Resources Nursery located in Xiuwen County, Guizhou Province. The plant species used in our studies are authenticated by Professor Baolin Guo. Specially, the plant selected for genome sequencing, as well as RNA-seq analysis across different tissues and leaf developmental stages, qRT-PCR and UPLC experimentation, was confirmed as E. pubescens . Additionally, Professor Baolin Guo identified the plants utilized in RNA-seq investigations of various flower hues, including E. pseudowushanense , E. acuminatum , E. jinchengshanense , E. baojingense and E. hunanense . Voucher specimens for all plant samples are preserved at the Plant Specimen Museum, part of the Institute of Medicinal Plant Resources Development, Chinese Academy of Medical Sciences (coded as IMD). The deposition numbers assigned to E. pubescens , E. pseudowushanense , E. acuminatum , E. jinchengshanense , E. baojingense and E. hunanense were B. L. Guo 0711-3, B. L. Guo 0312, B. L. Guo 0342, B. L. Guo 0524, B. L. Guo 0332 and B. L. Guo 0402, respectively.
PAL gene identification and sequence analysis
The g enome sequences of E. pubescens were accessed from the National Center for Biotechnology Information (NCBI) under project PRJNA747870 [ 35 ]. Utilizing APG IV [ 36 ] as a reference, genome sequences of 23 species were located and retrieved from Phytozome ( http://www.phytozome.net ) and the NCBI database ( https://www.ncbi.nlm.nih.gov/ ) (Table S1 and Table S2 ). The analyzed species are as follows: one algae ( Chara braunii ), one bryophyte ( Physcomitrella patens ), one fern ( Ceratopteris richardii ), two gymnosperms ( Ginkgo biloba and Sequoiadendron giganteum ), two basal angiosperms ( Amborella trichopoda and Nymphaea colorata ), four Magnoliidae ( Ceratophyllum demersum , Cinnamomum kanehirae , Liriodendron chinense , Piper nigrum ), a basal monocotyledon ( Spirodela polyrrhiza ), six early-diverging eudicotyledons ( Nelumbo nucifera , Macadamia integrifolia , Tetracentron sinense , Aquilegia coerulea , Kingdonia uniflora , Papaver somniferum , E. pubescens ), three typical dicotyledons ( Vitis vinifera , Cucumis sativus , Arabidopsis thaliana ), an early-diverging monocotyledons ( Brachypodium distachyon ) and a typical monocotyledon ( Oryza sativa ). Predicted proteins from these genomes underwent screening with HMMER v3 [ 37 ], employing the Hidden Markov Model (HMM) corresponding to Pfam [ 38 ] (PF00221; http://pfam.sanger.ac.uk/ ). Among the proteins identified using the PAL HMM, a subset of high-quality proteins (E-value < 1e-20 and confirmed integrity of the PAL domain) was selected for alignment. In cases where only genome information was available for certain species, a localized TBLASTN search was conducted against the PAL genes of A. thaliana and O. sativa , considering records with maximum identity > 95%, length > 400 bp, and E-value < 1e-20. To validate the results of the HMM and BLAST searches, all potential PAL genes were further subjected to analysis in the NCBI-CDD database ( https://www.ncbi.nlm.nih.gov/cdd/ ) to confirm the presence of conserved domains, and candidates lacking the “PAL-HAL” shorthand designation were discarded. Protein sequences were excluded if the PAL domain appeared truncated or if the PAL domain match E-value exceeded 1e-5. Following these stringent criteria, 167 PAL genes were ultimately identified across the nine species studied (Table S2 ). Sequences of the 7 EpPALs have been submitted to China National Center for Bioinformation (CNCB), with the accession numbers for EpPAL1 - EpPAL7 are C_AA071439.1, C_AA071440.1, C_AA071441.1, C_AA071442.1, C_AA071443.1, C_AA071444.1 and C_AA071445.1, respectively. The accession number of Ep-actin gene used in qPCR is C_AA071459.1.
Protein sequence properties analysis, conserved domain and motifs analysis
The physiological and biochemical characteristics of the full-length proteins were determined using the ProtParam tool ( http://web.expasy.org/protparam/ ) [ 39 ]. SignalP (V.4.1) ( http://www.cbs.dtu.dk/services/SignalP/ ) [ 40 ] and Euk-mPLoc (V.2.0) ( http://www.csbio.sjtu.edu.cn/bioinf/euk-multi-2/# ) [ 41 ] were utilized to analyze the signal peptide and subcellular localization of each protein, respectively. Additionally, MEME (V.5.0.2) ( http://meme-suite.org/ ) [ 42 ] was employed to identify conserved motifs, including the PAL domain, using optimized parameters: a maximum of 10 motifs were searched for, with each motif ranging from 6 to 50 residues in width.
Phylogenetic analysis, synteny block identification and gene duplication pattern analysis
The protein sequences were aligned using ClustalW2 [ 43 ] with its default settings. Phylogenetic trees were inferred using the maximum likelihood (ML) method with the JTT + R9 model, which was automatic selected by IQ-TREE [ 44 ]. The evolutionary tree was then visualized and further refined using iTOL ( https://itol.embl.de/ ) [ 45 ]. Synteny blocks between genomes and intra-specific collinearity analysis were identified using the jcvi pipeline ( https://github.com/tanghaibao/jcvi ). BLASTP was performed to identify paralogous or orthologous gene pairs, with an E-value cutoff of 1e-05. To identify patterns of gene duplication, DupGen-finder ( https://github.com/qiao-xin/DupGen_finder ) [ 46 ] was employed.
Chromosome location, cis -acting element and gene structure analysis
The gene location map was constructed using MapChart V.2.0 ( http://mg2c.iask.in/mg2c_v2.0/ ) [ 47 ]. Cis -acting elements located within the 1.5 kb upstream sequences of the 5′ regulatory region, starting from the transcriptional start site, were identified using PlantCARE ( http://bioinformatics.psb.ugent.be/webtools/plantcare/html/ ). To assess the divergence between upstream sequences of each paralogous gene pair, the GATA program [ 48 ] was employed with a window size of seven and a lower cut-off score of 12 bits. Lastly, the visualization of gene structure was facilitated by TBtools software [ 49 ].
Natural selection test
Codeml program in PAML (V.4.8) [ 50 ] was conducted to detect changes in evolutionary rates and signatures of positive selection. Four levels of positive selection tests were performed. (1) Detection of positive selection in pairwise genes of all EpPALs . For this, the main parameter settings were: runmode = -2 and NSsites = 0; (2) Site-specific model was applied for positive selection detection of sites in genes. This model assumes a constant ω (ω = dN/dS; where dN is the non-synonymous substitution rate and dS is the synonymous substitution rate) across all branches. The main parameters were set as runmode = 0 and NSites = 0 1 2 7 8. To determine the most suitable model for detection, we compared Neutral M1 vs. Selection M1 and M7 vs. M8; (3) Branch model was applied to detect the rapidly evolving genes in the target branch. This model assumes a constant ω for all sites with a gene. Three scenarios were tested: the one-ratio model (assuming a constant ω for all branches with parameters as model = 0 and NSites = 0), the two-ratio model (assuming a foreground ω for designated branches and a background ω for all others with parameters as model = 2 and Nsites = 0), and the free-ratio model (allowing different ω for each branch with parameters as model = 1 and Nsites = 0) [ 51 ]. Models were compared using likelihood ratio tests based on the log likelihood (lnL). The chi-square test with a significance threshold of P < 0.05 was used to compare 2|ΔlnL| values between models; (4) Branch-site model was applied to detect whether there exist positive selection sites in a specific branch. This model assumes one ω for the target branch and another constant ω for all other branches. We compared the branch-site model A (model = 2, NSites = 2, fix_omega = 0, omega = 2) with its null model (model = 2, NSites = 2, fix_omega = 1, omega = 1). If the chi-square test yielded a significance of P < 0.05, we employed the Bayes Empirical Bayes (BEB) method to calculate the posterior probability. Genes in the specific branch were considered under positive selection if this value exceeded 0.95 [ 52 ].
RNA-seq and correlation analysis
To identify the gene expression profiles of EpPALs , we conducted three independent RNA-seq experiments. Firstly, we sampled different tissues (roots, stems, leaves, flowers, and fruits) from E. pubescens . Secondly, we collected leaves from various developmental stages of E. pubescens , specifically: Stage 1 (S1) with leaf width of 0.5–1 cm and low leatheriness; Stage 2 (S2) with leaf width of 1.5–2 cm and low leatheriness; Stage 3 (S3) with leaf width of 2.5 ~ 4 cm and low leatheriness; Stage 4 (S4) with leaf width of 5 cm and medium leatheriness; and Stage 5 (S5) with leaf width of 5 cm and high leatheriness. Thirdly, we included six species of Epimedium with diverse petal colors (magenta in E. pseudowushanense and E. acuminatum , yellow in E. jinchengshanense , and green in E. baojingense ), sepal colors (red in E. hunanense , green in E. baojingense , and white in E. acuminatum ), and leaf colors (green and magenta in E. pubescens ). The RNA-seq protocol and classification criteria followed Xu et al. (2023) [ 53 ]. Initial protein contamination screening was performed using the NanoDrop ND 1000 (Nanodrop technologies), ensuring a tightly controlled OD260/OD280 ratio within the range of 1.9 to 2.1. Subsequently, the RNA Integrity Number (RIN) was evaluated using the Agilent Technologies 2100 bioanalyzer (Agilent, Santa Clara, CA). Sequencing was only initiated if the RIN exceeded 8 and the 28 S/18S ratio was greater than or equal to 0.7. Software tools including Trimmomatic (version 0.36) [ 54 ], HISAT2 [ 55 ], and the R package Rsubread [ 56 ] were utilized for quality control, sequence alignments, and gene expression quantification, respectively. The reference genome utilized was that of E. pubescens , as published by Shen et al. (2022) [ 35 ]. All samples were collected between 10:00–11:30 am on a sunny day and immediately treated with liquid nitrogen before being stored in dry ice for transport to Beijing. All samples were conserved at -80 °C under ultralow temperature for subsequent RNA extraction and chemical component identification. We used the R packages Tidyverse and ggcor to compute the pearson correlation between PALs and the relative content of PFGs.
UPLC experiment
For the analysis of PFGs content, approximately 0.1 g of ground sample was soaked in 10 ml of 50% ethanol and ultrasonicated for 30 min before being filtrated through 0.22 μm filter membrane (Millipore, Nylon) for UPLC analysis. UPLC under 270 nm was conducted at a flow rate of 0.3 ml/min using the ACQUITY UPLC system (UPLC I-class; Waters, Milford, MA, USA) equipped with an ACQUITY UPLC BEH C18 column (2.1 × 100 mm. 1.7 μm; Waters, Milford, MA, USA) maintained at 25 °C. The mobile phase comprised of water (eluent A) and 100% acetonitrile (eluent B). The authentic flavonoids were purchased from the Shanghai Yuanye Bio-Technology Co., Ltd., Shanghai, China.
To ensure the reliability of our transcriptome data, we conducted qRT-PCR analysis on leaves from five distinct developmental stages of E. pubescens , focusing on six selected EpPALs . Total RNA was extracted using a plant total RNA extraction kit from Aidlab (China). We assessed RNA integrity on a 1.2% agarose gel and quantified it using a NanoDrop 2000 C Spectrophotometer from Thermo Scientific (USA). cDNA synthesis was achieved using the TransScript One-Step gDNA Removal and cDNA Synthesis SuperMix Kit from Transgen Biotech (China). qRT-PCR reactions were performed for each tissue sample with gene-specific primers (Table S10 ). The qRT-PCR program consisted of pre-denaturation at 95 °C for 2 min, followed by 40 cycles of amplification at 95 °C for 15 s, 60 °C for 30 s, and 72 °C for 30 s. We analyzed the relative abundance of transcripts using the comparative Ct method, applying the formula 2-ΔΔCt for relative quantification. Our gene expression results were calculated based on the 2-ΔΔCt method, and the reported data represents the average of three biological and three technical replicates.
7 PALs were firstly and comprehensively identified based on the genome of E. pubescens . EpPAL2 , EpPAL3 and EpPAL1 were identified as the ancient isoforms. EpPAL2 and EpPAL3 exhibited a homology range of 61.09 to 64.38%, contained two introns and underwent strong purifying selection, evolving at a rate ~ 10 to ~ 54 times slower compared to EpPAL1 and modern EpPALs ( EpPAL4-7 ). The evolutionary trajectory of modern EpPALs was shaped by multiple duplication events. Initially, EpPAL4 emerged through intraspecific whole-genome duplication from EpPAL1 . This was followed by a sequence of tandem duplications resulting in EpPAL5 and EpPAL6 , and transposed duplications that gave rise to EpPAL7 , all originating from EpPAL4 . Analysis of expression profiles through RNA-seq and UPLC techniques revealed that EpPAL2 and EpPAL3 are key genes involved in the biosynthesis of prenylated flavonol glycosides. This finding was further validated through parallel UPLC and qRT-PCR experiments. Novel insights into the evolution of 24 PAL gene families were provided, revealing the evolutionary characteristics of 12 different evolutionary clade groups. Overall, this study offers a unique perspective on PAL evolution and clarifies the role of PAL genes in Epimedium plants.
Data availability
The experimental materials were stored at the Institute of Medicinal Plant Development, Chinese Academy of Medical Sciences. The datasets containing the E. pubescens genome sequences can be accessed from the National Center for Biotechnology Information (NCBI) repository using the accession number PRJNA747870. Additionally, the RNA-seq datasets generated in this study have been deposited in the China National Center for Bioinformation (CNCB) repository. Specifically, the RNA-seq data related to different tissues of E. pubescens, various developmental stages of E. pubescens leaves, and six species of Epimedium with distinct petal colors can be retrieved using the accession numbers CRA014527, CRA014549, and CRA014550, respectively.
Naoumkina M, Zhao Q, Gallego-Giraldo L, Dai X, Zhao PX, Dixon R. Genome-wide analysis of phenylpropanoid defence pathways. Mol Plant Pathol. 2010;11(6):829–46.
Article CAS PubMed PubMed Central Google Scholar
Barros J, Dixon RA. Plant phenylalanine/tyrosine ammonia-lyases. Trends Plant Sci. 2020;25(1):66–79.
Article CAS PubMed Google Scholar
Bennici A. Origin and early evolution of land plants: problems and considerations. Commun Integr Biol. 2008;1(2):212–8.
Article PubMed PubMed Central Google Scholar
Shalaby S, Horwitz BA. Plant phenolic compounds and oxidative stress: integrated signals in fungal-plant interactions. Curr Genet. 2015;61(3):347–57.
Liu CW, Murray JD. The role of flavonoids in nodulation host-range specificity: an update. Plants (Basel). 2016;5(3):33.
MacDonald MJ, D’Cunha GB. A modern view of phenylalanine ammonia lyase. Biochem Cell Biol. 2007;85(3):273–82.
Schwede TF, Rétey J, Schulz GE. Crystal structure of histidine ammonia-lyase revealing a novel polypeptide modification as the catalytic electrophile. Biochemistry. 1999;38(17):5355–61.
Dong C, Cao N, Zhang Z, Shang Q. Phenylalanine ammonia-lyase gene families in cucurbit species: structure, evolution, and expression. J Integr Agric. 2016;15(6):1239–55.
Article CAS Google Scholar
Chang A, Lim MH, Lee SW, Robb EJ, Nazar RN. Tomato phenylalanine ammonia-lyase gene family, highly redundant but strongly underutilized. J Biol Chem. 2008;283(48):33591–601.
Bate NJ, Orr J, Ni W, Meromi A, Nadler-Hassar T, Doerner PW, et al. Quantitative relationship between phenylalanine ammonia-lyase levels and phenylpropanoid accumulation in transgenic tobacco identifies a rate-determining step in natural product synthesis. J Clin Periodontol. 1994;91(16):7608–12.
CAS Google Scholar
Huang J, Gu M, Lai Z, Fan B, Shi K, Zhou YH, et al. Functional analysis of the arabidopsis PAL gene family in plant growth, development, and response to environmental stress. Plant Physiol. 2010;153(4):1526–38.
de Jong F, Hanley SJ, Beale MH, Karp A. Characterisation of the willow phenylalanine ammonia-lyase (PAL) gene family reveals expression differences compared with poplar. Phytochemistry. 2015;117:90–7.
Ma H, He X, Yang Y, Li M, Hao D, Jia Z. The genus Epimedium : an ethnopharmacological and phytochemical review. J Ethnopharmacol. 2011;134(3):519–41.
Jiang J, Zhao Bj, Song J, Jia X. Pharmacology and clinical application of plants in Epimedium L. Chin Herb Med. 2016;8(1):12–23.
Zhu Jf L, Zj Z, Gs, Meng K, Kuang Wy, Li J, et al. Icaritin shows potent anti-leukemia activity on chronic myeloid leukemia in vitro and in vivo by regulating MAPK/ERK/JNK and JAK2/STAT3 /AKT signalings. PLoS ONE. 2011;6(8):e23720.
Zhao H, Guo Y, Li S, Han R, Ying J, Zhu H, et al. A novel anti-cancer agent Icaritin suppresses hepatocellular carcinoma initiation and malignant growth through the IL-6/Jak2/Stat3 pathway. Oncotarget. 2015;6(31):31927–43.
Zeng S, Liu Y, Zou C, Huang W, Wang Y. Cloning and characterization of phenylalanine ammonia-lyase in medicinal Epimedium species. Plant Cell Tissue Organ Cult. 2013;113(2):257–67.
Liu Y, Wu L, Deng Z, Yu Y. Two putative parallel pathways for naringenin biosynthesis in Epimedium wushanense . RSC Adv. 2021;11(23):13919–27.
Wu Z, Gui S, Wang S, Ding Y. Molecular evolution and functional characterisation of an ancient phenylalanine ammonia-lyase gene (NnPAL1) from Nelumbo nucifera : novel insight into the evolution of the PAL family in angiosperms. BMC Evol Biol. 2014;14:100.
Dong CJ, Shang QM. Genome-wide characterization of phenylalanine ammonia-lyase gene family in watermelon ( Citrullus lanatus ). Planta. 2013;238(1):35–49.
Li G, Wang H, Cheng X, Su X, Zhao Y, Jiang T, et al. Comparative genomic analysis of the PAL genes in five Rosaceae species and functional identification of Chinese white pear. PeerJ. 2019;7:e8064.
Yan F, Li H, Zhao P. Genome-wide identification and transcriptional expression of the PAL gene family in common walnut ( Juglans Regia L). Genes. 2019;10(1):46.
Hou X, Shao F, Ma Y, Lu S. The phenylalanine ammonia-lyase gene family in Salvia miltiorrhiza : genome-wide characterization, molecular cloning and expression analysis. Mol Biol Rep. 2013;40(7):4301–10.
Thiyagarajan K, Vitali F, Tolaini V, Galeffi P, Cantale C, Vikram P, et al. Genomic characterization of phenylalanine ammonia lyase gene in buckwheat. PLoS ONE. 2016;11(3):e0151187.
Bagal UR, Leebens-Mack JH, Lorenz WW, Dean JFD. The phenylalanine ammonia lyase (PAL) gene family shows a gymnosperm-specific lineage. BMC Genomics. 2012;13(3):S1.
Duret L. Why do genes have introns? Recombination might add a new piece to the puzzle. Trends Genet. 2001;17(4):172–5.
Wang HF, Feng L, Niu DK. Relationship between mRNA stability and intron presence. Biochem Biophys Res Commun. 2007;354(1):203–8.
Vogt T. Phenylpropanoid biosynthesis. Mol Plant. 2010;3(1):2–20.
Hu GS, Jia JM, Hur YJ, Chung YS, Lee JH, Yun DJ, et al. Molecular characterization of phenylalanine ammonia lyase gene from Cistanche deserticola . Mol Biol Rep. 2011;38(6):3741–50.
Wanner LA, Li G, Ware D, Somssich IE, Davis KR. The phenylalanine ammonia-lyase gene family in Arabidopsis thaliana . Plant Mol Biol. 1995;27(2):327–38.
Olsen KM, Lea US, Slimestad R, Verheul M, Lillo C. Differential expression of four Arabidopsis PAL genes; PAL1 and PAL2 have functional specialization in abiotic environmental-triggered flavonoid synthesis. J Plant Physiol. 2008;165(14):1491–9.
Habibollahi M, Kavousi HR, Lohrasbi-Nejad A, Rahpeyma SA. Cloning, characterization and expression of a phenylalanine ammonia-lyase gene ( CcPAL ) from cumin ( Cuminum cyminum L). J Appl Res Med Aromat Plants. 2020;18:100253.
Google Scholar
Duarte JM, Cui L, Wall PK, Zhang Q, Zhang X, Leebens-Mack J, et al. Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis . Mol Biol Evol. 2006;23(2):469–78.
Lei L, Zhou SL, Ma H, Zhang LS. Expansion and diversification of the SET domain gene family following whole-genome duplications in Populus trichocarpa . BMC Evol Biol. 2012;12:51.
Shen G, Luo Y, Yao Y, Meng G, Zhang Y, Wang Y, et al. The discovery of a key prenyltransferase gene assisted by a chromosome-level Epimedium pubescens genome. Front Plant Sci. 2022;13:1034943.
Group TAP, Chase MW, Christenhusz MJM, Fay MF, Byng JW, Judd WS, et al. An update of the angiosperm phylogeny group classification for the orders and families of flowering plants: APG IV. Bot J Linn Soc. 2016;181(1):1–20.
Article Google Scholar
Finn RD, Clements J, Eddy SR. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 2011; 39(Web Server issue):W29–37.
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, et al. Pfam: the protein families database. Nucleic Acids Res. 2014;42(Database issue):D222–230.
Wang L, Wang L, Zhang Z, Ma M, Wang R, Qian M, et al. Genome-wide identification and comparative analysis of the superoxide dismutase gene family in pear and their functions during fruit ripening. Postharvest Biol Technol. 2018;143:68–77.
Petersen TN, Brunak S, von Heijne G, Nielsen H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods. 2011;8(10):785–6.
Chou KC, Shen HB. A new method for predicting the subcellular localization of eukaryotic proteins with both single and multiple sites: Euk-mPLoc 2.0. PLoS ONE. 2010;5(4):e9931.
Bailey TL, Johnson J, Grant CE, Noble WS. The MEME suite. Nucleic Acids Res. 2015;43(W1):W39–49.
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, et al. Clustal W and Clustal X version 2.0. Bioinformatics. 2007;23(21):2947–8.
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. 2020;37(5):1530–4.
Letunic I, Bork P. Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. 2019;47(W1):W256–9.
Qiao X, Li Q, Yin H, Qi K, Li L, Wang R, et al. Gene duplication and evolution in recurring polyploidization-diploidization cycles in plants. Genome Biol. 2019;20(1):38.
Voorrips RE. MapChart: software for the graphical presentation of linkage maps and QTLs. J Heredity. 2002;93(1):77–8.
Nix D, Eisen M. GATA: a graphic alignment tool for comparative sequence analysis. BMC Bioinformatics. 2005;6:9.
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant. 2020;13(8):1194–202.
Yang Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol. 2007;24(8):1586–91.
Yang Z. Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol. 1998;15(5):568–73.
Yang Z, Nielsen R. Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol. 2002;19(6):908–17.
Xu C, Liu X, Shen G, Fan X, Zhang Y, Sun C, et al. Time-series transcriptome provides insights into the gene regulation network involved in the icariin-flavonoid metabolism during the leaf development of Epimedium pubescens . Front Plant Sci. 2023;14:1183481.
Anthony M, Marc L, Bjoern U. Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics. 2014;30(15):2114–20.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Liao Y, Smyth GK, Shi W. The r package rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nuc Acids Res. 2019;47(8):e47.
Download references
Acknowledgements
We thank all our colleagues for providing useful discussions and technical assistance. We are very grateful to the editor and reviewers for critically evaluating the manuscript and providing constructive comments for its improvement.
This work was financially supported by the CAMS Innovation Fund for Medical Sciences (CIFMS) (2021-I2M-1-031).
Author information
Chaoqun Xu and Xuelan Fan contributed equally to this work and share the first authorship.
Authors and Affiliations
Key Laboratory of Bioactive Substances and Resources Utilization of Chinese Herbal Medicines, Ministry of Education, Institute of Medicinal Plant Development, Peking Union Medical College and Chinese Academy of Medical Sciences, No.151 MaLianWa North Road, Haidian District, Beijing, 100193, China
Chaoqun Xu, Xuelan Fan, Guoan Shen & Baolin Guo
College of Pharmacy, Jiangxi University of Chinese Medicine, Nanchang, 330004, China
You can also search for this author in PubMed Google Scholar
Contributions
Chaoqun Xu conceived and designed the study, put into effect the main bioinformatics analyses, wrote the manuscript, and prepared the figures and tables. Xuelan Fan prepared the materials, conducted the experiments, data analysis, and revised the manuscript drafts. Guoan Shen participated in the design of this study and revised the manuscript. Baolin Guo conceived and designed the study, involved in data interpretation and finalizing the manuscript draft. The authors read and approved the final manuscript.
Corresponding author
Correspondence to Baolin Guo .
Ethics declarations
Ethics approval and consent to participate.
All the experimental materials in this study have obtained the authority. The experimental research and method on all the experimental materials, including the collection of plant material, comply with relevant institutional, national, and international guidelines.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1
Supplementary material 2, supplementary material 3, supplementary material 4, supplementary material 5, supplementary material 6, rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/ .
Reprints and permissions
About this article
Cite this article.
Xu, C., Fan, X., Shen, G. et al. Genome-wide identification of the phenylalanine ammonia-lyase gene from Epimedium Pubescens Maxim. (Berberidaceae): novel insight into the evolution of the PAL gene family. BMC Plant Biol 24 , 831 (2024). https://doi.org/10.1186/s12870-024-05480-z
Download citation
Received : 09 January 2024
Accepted : 01 August 2024
Published : 04 September 2024
DOI : https://doi.org/10.1186/s12870-024-05480-z
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Epimedium pubescens
- Phenylalanine ammonia-lyase gene
- Prenylated flavonol glycoside
- Expression profiling
BMC Plant Biology
ISSN: 1471-2229
- General enquiries: [email protected]

IMAGES
VIDEO
COMMENTS
In brief: What types of studies are there? - InformedHealth.org
Types of Research - Explained with Examples
6 Basic Types of Research Studies (Plus Pros and Cons)
Types of Research Designs Compared | Guide & Examples
Types of Research Studies. ... Click on each type of research study to see a research article example. ... an experimental design in which multiple measures are collected over a period of time from two or more groups of different ages (birth cohorts), ethnicities, or other factos. These designs combine aspects of longitudinal design and cohort ...
Study designs: Part 1 - An overview and classification - PMC
What are some different types of research studies?
Types of studies and research design - PMC
The Two Main Research Types - A Brief Overview
Evidence-Based Medicine: Types of Studies
Qualitative vs. Quantitative Research | Differences, ...
What Are the Different Types of Clinical Research?
A meta-analysis is a type of systematic review that goes one step further, combining the data from multiple studies and using statistics to summarize it, as if creating a mega-study from many smaller studies.4. However, even systematic reviews and meta-analyses aren't the final word on scientific questions.
What Is a Research Design | Types, Guide & ...
Understanding Research Study Designs - PMC
Cohort Studies. These are observational studies that follow large groups of people over a long period of time, years or even decades, to find associations of an exposure (s) with disease outcomes. Researchers regularly gather information from the people in the study on several variables (like meat intake, physical activity level, and weight).
Types of Research Methods. Research methods can be broadly categorized into two types: quantitative and qualitative. Quantitative methods involve systematic empirical investigation of observable phenomena via statistical, mathematical, or computational techniques, providing an in-depth understanding of a specific concept or phenomenon (Schweigert, 2021).
19 Types of Research (With Definitions and Examples)
Types of Research Studies; Search this Guide Search. Research, Statistics, Writing, Dissertation & Publishing. ... an experimental design in which multiple measures are collected over a period of time from two or more groups of different ages (birth cohorts), ethnicities, or other factos. These designs combine aspects of longitudinal design and ...
Epidemiology is defined as the study of human populations. These studies often investigate the relationship between dietary consumption and disease development. There are three main types of epidemiological studies: cross-sectional, case-control, and prospective cohort studies. Figure 2.2: Types of epidemiology.
Clinical research study designs: The essentials - PMC
Research Methods | Definitions, Types, Examples
Human-bird conflicts are in a critical state, involving economic losses such as agricultural losses, bird strikes on aircraft and avian influenza. Traditional technologies leveraging bird vision and hearing often lose their effectiveness over time as birds become habituated to these stimuli. To address these challenges, our study introduces a novel countermeasure technology based on ...
From among Charlotte Bühler's (1893-1974) contributions to child-psychology, psychology of adolescence, life-span, and later humanistic psychology, this paper deals with experiments conducted by her team in the Twenties in Vienna, and recorded in two articles on "Affective Effects of Impressions of Strangeness in the first year of life" and "Two Main Types of Life Processes", neither ...
Types of Study in Medical Research - PMC
Significance tests for each EpPAL gene in two types of leaves were conducted and were labled with italicized ... Previous studies have primarily focused on research at the species or family level [20, 21, 23, 24, 29], with limited investigations on the evolution of PAL genes, except for those in N. nucifera and cucurbit species .